orthodb
1、数据库

orthodb数据:
odb10v0_levels.tab.gz: NCBI taxonomy nodes where Ortho DB orthologous groups (OGs) are calculated
odb10v0_species.tab.gz: Ortho DB individual organism (aka species) ids based on NCBI taxonomy ids (mostly species level)
odb10v0_level2species.tab.gz: correspondence between level ids and species ids
odb10v0_genes.tab.gz: Ortho DB genes with some info
odb10v0_OGs.tab.gz: Ortho DB orthologous groups
odb10v0_OG2genes.tab.gz: OGs to genes correspondence
odb10v0_OG_xrefs.tab.gz: OG associations with GO, COG and InterPro ids
v9_v10_OGs_map.tab.gz mappings between the previous and current release orthologous group ids
odb10v0_fasta_<root>.tgz tar-ball with one fasta file per taxon id in the given root (bacteria,metazoa,fungi,plants)
2、odb10v0_levels.tab:
1. level NCBI taxonomy id
2. scientific name
3. total non-redundant count of genes in all underneath clustered species(在聚集的物种下面的所有的基因的总非重复计数)
4. total count of OGs built on it
5. total non-redundant count of species underneath

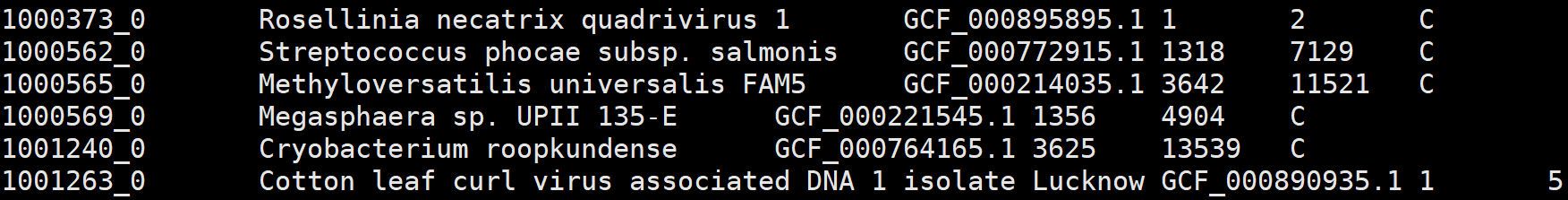
3、odb10v0_species.tab.gz
1. Ortho DB individual organism id, based on NCBI tax id
2. scientific name inherited from the most relevant NCBI tax id
3. genome asssembly id, when available
4. total count of clustered genes in this species
5. total count of the OGs it participates
6. mapping type, clustered(C) or mapped(M)
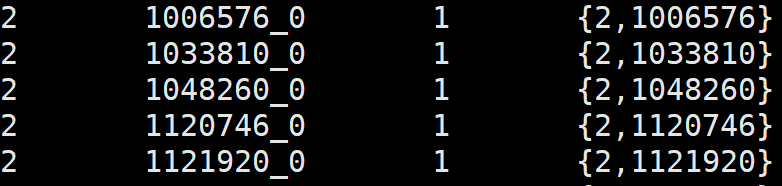
4、odb10v0_level2species.tab
1. top-most level NCBI tax id, one of [2,2157,2759,10239]
2. Ortho DB organism id
3. number of hops between the top-most level id and the NCBI tax id assiciated with the organism
4. ordered list of Ortho DB selected intermediate levels from the top-most level to the bottom one
5、odb10v0_genes.tab
1. Ortho DB unique gene id (not stable between releases)
2. organism tax id
3. protein original sequence id, as downloaded along with the sequence
4. Uniprot id, evaluated by mapping
5. ENSEMBL gene name, evaluated by mapping
6. NCBI gid, evaluated by mapping
7. description, evaluated by mapping
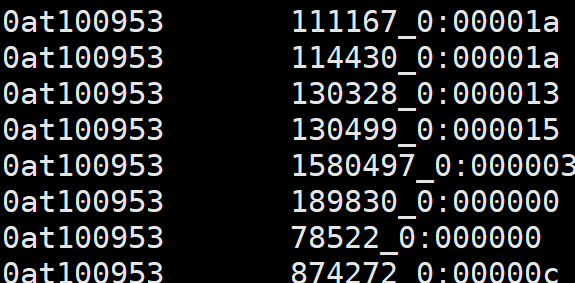
6、odb10v0_OG2genes.tab
1. OG unique id
2. Ortho DB gene id
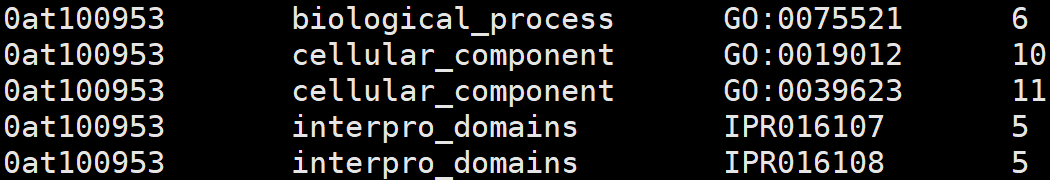
7、odb10v0_OG_xrefs.tab
1. OG unique id
2. external DB or DB section
3. external identifier
4. number of genes in the OG associated with the identifier
参考
https://www.orthodb.org/?page=filelist
orthodb的更多相关文章
- 【基因组预测】braker2基因结构注释要点记录
目录 流程使用 问题 记录下braker2的使用要点,以备忘记. 流程使用 braker2有很多流程,根据你的数据:组装的基因组.转录组.蛋白(同源,包括近缘或远缘)选择不同流程,官网有说明: htt ...
随机推荐
- failed to open stream: Permission denied in警告错误
问题是文件所在目录的权限问题导致的.只需要将警告文件所在的目录权限更改为777(至少是006)即可 例如 (...a.log)failed to open stream: Permission den ...
- UVA-10020-贪心
题意:给你一些数轴上的线段,要求寻找出某些线段能够完全覆盖[0,M],并且取的线段数目最小. 解题思路: 贪心思路, 1.每个线段都有一个L和R,代表它的起点和终点,对于所有R <= 0 , ...
- HDFS 异构储存配置及基本命令操作
hadoop-2.8.4 部署我就不说了 网上一大堆 hdfs-site.xml datanode 储存路径挂载需要修改如下: <property> <name>dfs.dat ...
- hive命令的执行方式
1.通过cli直接执行 2.hive -e "hql" 如:[root@host ~]# hive -e "use gamedw;show tables" [r ...
- elcipse 安装lombok插件解决 @Slf4j 等找不到log变量问题
参考:http://blog.51cto.com/4925054/2127840 <dependency> <groupId>org.projectlombok</gro ...
- Intellij IDEA编辑golang时无法加载系统GOPATH变量
问题: 编译go项目时,报找不到包.从日志看,GOPATH与系统设置的不一致. 如何解决:系统的gopath路径,加到Project libraries中 参考:https://segmentfaul ...
- WebService与RESTful WebService
Manual Instruction Document Web Service JAX-WS & JAX-RS Author: Liu Xiang Date: 2018/01/12 1. Su ...
- ABAP-串口通信-道闸设备
最近SAP系统需要与道闸设备集成,通过串口通讯模式控制道闸栏杆升降,在此将开发过程中的思路及问题点做个备注. 一.相关设备 道闸设备型号:富士智能FJC-D618 串口模块:康耐德 C2000-A1- ...
- ubuntu交换Caps 和 ESC
https://askubuntu.com/questions/363346/how-to-permanently-switch-caps-lock-and-esc This will allow y ...
- 尚硅谷redis学习3-redis启动以后的杂项
redis速度很快,运行benchmark可以看出,各项运行速度可达100000次每秒 redis默认有16个数据库,分别是0, 1 ... 15,默认在0号库,可以通过select num转到其它库 ...
