[占位-未完成]scikit-learn一般实例之十二:用于RBF核的显式特征映射逼近
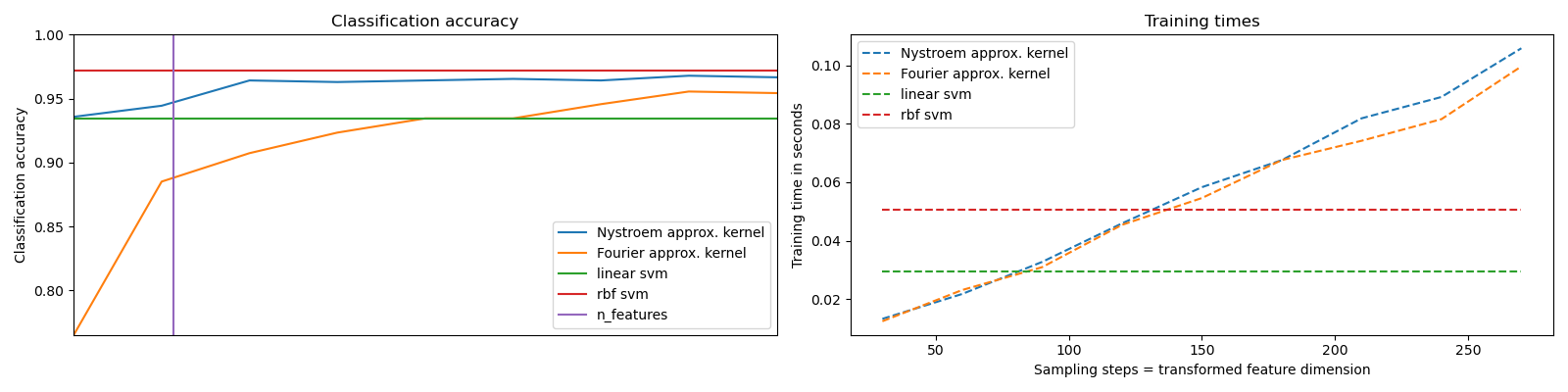
It shows how to use RBFSampler and Nystroem to approximate the feature map of an RBF kernel for classification with an SVM on the digits dataset. Results using a linear SVM in the original space, a linear SVM using the approximate mappings and using a kernelized SVM are compared. Timings and accuracy for varying amounts of Monte Carlo samplings (in the case of RBFSampler, which uses random Fourier features) and different sized subsets of the training set (for Nystroem) for the approximate mapping are shown.
Please note that the dataset here is not large enough to show the benefits of kernel approximation, as the exact SVM is still reasonably fast.
Sampling more dimensions clearly leads to better classification results, but comes at a greater cost. This means there is a tradeoff between runtime and accuracy, given by the parameter n_components. Note that solving the Linear SVM and also the approximate kernel SVM could be greatly accelerated by using stochastic gradient descent via sklearn.linear_model.SGDClassifier. This is not easily possible for the case of the kernelized SVM.
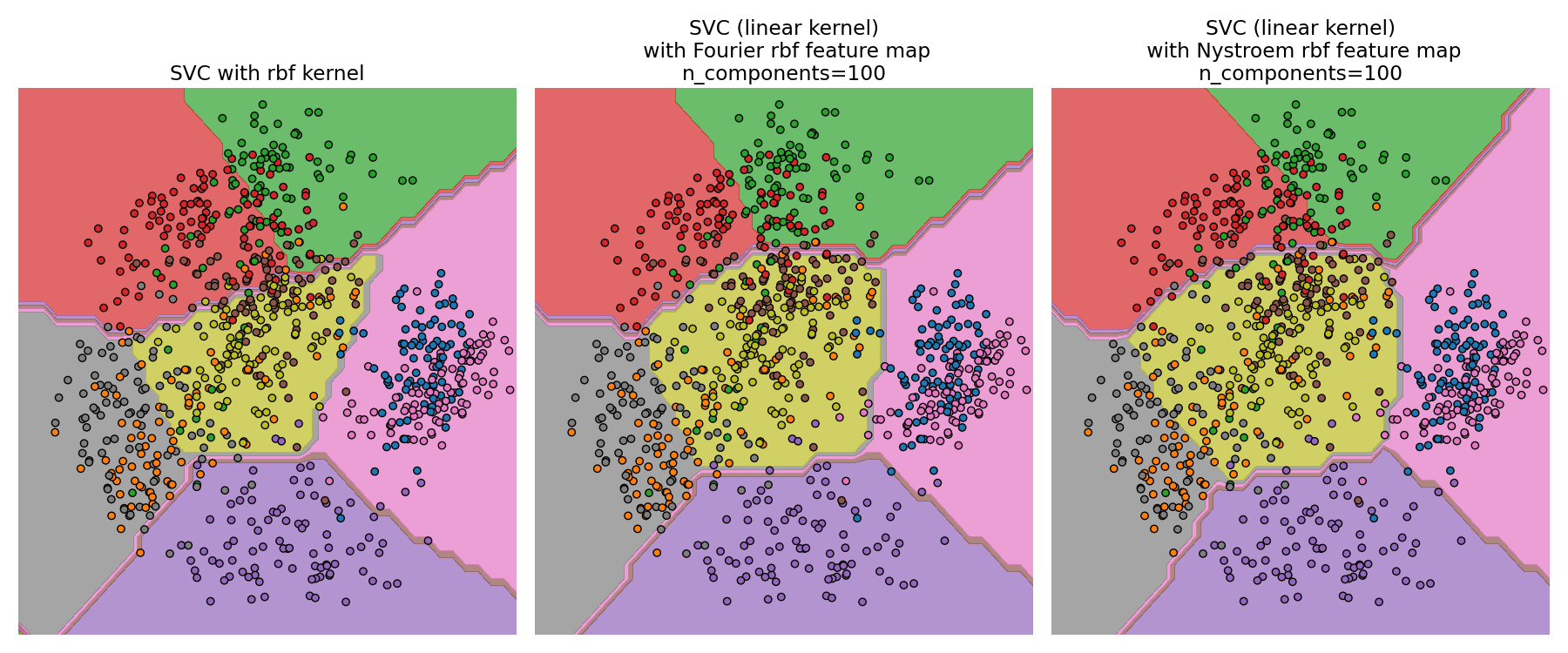
The second plot visualized the decision surfaces of the RBF kernel SVM and the linear SVM with approximate kernel maps. The plot shows decision surfaces of the classifiers projected onto the first two principal components of the data. This visualization should be taken with a grain of salt since it is just an interesting slice through the decision surface in 64 dimensions. In particular note that a datapoint (represented as a dot) does not necessarily be classified into the region it is lying in, since it will not lie on the plane that the first two principal components span.
The usage of RBFSampler and Nystroem is described in detail in Kernel Approximation.
print(__doc__)
# Author: Gael Varoquaux <gael dot varoquaux at normalesup dot org>
# Andreas Mueller <amueller@ais.uni-bonn.de>
# License: BSD 3 clause
# Standard scientific Python imports
import matplotlib.pyplot as plt
import numpy as np
from time import time
# Import datasets, classifiers and performance metrics
from sklearn import datasets, svm, pipeline
from sklearn.kernel_approximation import (RBFSampler,
Nystroem)
from sklearn.decomposition import PCA
# The digits dataset
digits = datasets.load_digits(n_class=9)
# To apply an classifier on this data, we need to flatten the image, to
# turn the data in a (samples, feature) matrix:
n_samples = len(digits.data)
data = digits.data / 16.
data -= data.mean(axis=0)
# We learn the digits on the first half of the digits
data_train, targets_train = data[:n_samples / 2], digits.target[:n_samples / 2]
# Now predict the value of the digit on the second half:
data_test, targets_test = data[n_samples / 2:], digits.target[n_samples / 2:]
#data_test = scaler.transform(data_test)
# Create a classifier: a support vector classifier
kernel_svm = svm.SVC(gamma=.2)
linear_svm = svm.LinearSVC()
# create pipeline from kernel approximation
# and linear svm
feature_map_fourier = RBFSampler(gamma=.2, random_state=1)
feature_map_nystroem = Nystroem(gamma=.2, random_state=1)
fourier_approx_svm = pipeline.Pipeline([("feature_map", feature_map_fourier),
("svm", svm.LinearSVC())])
nystroem_approx_svm = pipeline.Pipeline([("feature_map", feature_map_nystroem),
("svm", svm.LinearSVC())])
# fit and predict using linear and kernel svm:
kernel_svm_time = time()
kernel_svm.fit(data_train, targets_train)
kernel_svm_score = kernel_svm.score(data_test, targets_test)
kernel_svm_time = time() - kernel_svm_time
linear_svm_time = time()
linear_svm.fit(data_train, targets_train)
linear_svm_score = linear_svm.score(data_test, targets_test)
linear_svm_time = time() - linear_svm_time
sample_sizes = 30 * np.arange(1, 10)
fourier_scores = []
nystroem_scores = []
fourier_times = []
nystroem_times = []
for D in sample_sizes:
fourier_approx_svm.set_params(feature_map__n_components=D)
nystroem_approx_svm.set_params(feature_map__n_components=D)
start = time()
nystroem_approx_svm.fit(data_train, targets_train)
nystroem_times.append(time() - start)
start = time()
fourier_approx_svm.fit(data_train, targets_train)
fourier_times.append(time() - start)
fourier_score = fourier_approx_svm.score(data_test, targets_test)
nystroem_score = nystroem_approx_svm.score(data_test, targets_test)
nystroem_scores.append(nystroem_score)
fourier_scores.append(fourier_score)
# plot the results:
plt.figure(figsize=(8, 8))
accuracy = plt.subplot(211)
# second y axis for timeings
timescale = plt.subplot(212)
accuracy.plot(sample_sizes, nystroem_scores, label="Nystroem approx. kernel")
timescale.plot(sample_sizes, nystroem_times, '--',
label='Nystroem approx. kernel')
accuracy.plot(sample_sizes, fourier_scores, label="Fourier approx. kernel")
timescale.plot(sample_sizes, fourier_times, '--',
label='Fourier approx. kernel')
# horizontal lines for exact rbf and linear kernels:
accuracy.plot([sample_sizes[0], sample_sizes[-1]],
[linear_svm_score, linear_svm_score], label="linear svm")
timescale.plot([sample_sizes[0], sample_sizes[-1]],
[linear_svm_time, linear_svm_time], '--', label='linear svm')
accuracy.plot([sample_sizes[0], sample_sizes[-1]],
[kernel_svm_score, kernel_svm_score], label="rbf svm")
timescale.plot([sample_sizes[0], sample_sizes[-1]],
[kernel_svm_time, kernel_svm_time], '--', label='rbf svm')
# vertical line for dataset dimensionality = 64
accuracy.plot([64, 64], [0.7, 1], label="n_features")
# legends and labels
accuracy.set_title("Classification accuracy")
timescale.set_title("Training times")
accuracy.set_xlim(sample_sizes[0], sample_sizes[-1])
accuracy.set_xticks(())
accuracy.set_ylim(np.min(fourier_scores), 1)
timescale.set_xlabel("Sampling steps = transformed feature dimension")
accuracy.set_ylabel("Classification accuracy")
timescale.set_ylabel("Training time in seconds")
accuracy.legend(loc='best')
timescale.legend(loc='best')
# visualize the decision surface, projected down to the first
# two principal components of the dataset
pca = PCA(n_components=8).fit(data_train)
X = pca.transform(data_train)
# Generate grid along first two principal components
multiples = np.arange(-2, 2, 0.1)
# steps along first component
first = multiples[:, np.newaxis] * pca.components_[0, :]
# steps along second component
second = multiples[:, np.newaxis] * pca.components_[1, :]
# combine
grid = first[np.newaxis, :, :] + second[:, np.newaxis, :]
flat_grid = grid.reshape(-1, data.shape[1])
# title for the plots
titles = ['SVC with rbf kernel',
'SVC (linear kernel)\n with Fourier rbf feature map\n'
'n_components=100',
'SVC (linear kernel)\n with Nystroem rbf feature map\n'
'n_components=100']
plt.tight_layout()
plt.figure(figsize=(12, 5))
# predict and plot
for i, clf in enumerate((kernel_svm, nystroem_approx_svm,
fourier_approx_svm)):
# Plot the decision boundary. For that, we will assign a color to each
# point in the mesh [x_min, x_max]x[y_min, y_max].
plt.subplot(1, 3, i + 1)
Z = clf.predict(flat_grid)
# Put the result into a color plot
Z = Z.reshape(grid.shape[:-1])
plt.contourf(multiples, multiples, Z, cmap=plt.cm.Paired)
plt.axis('off')
# Plot also the training points
plt.scatter(X[:, 0], X[:, 1], c=targets_train, cmap=plt.cm.Paired)
plt.title(titles[i])
plt.tight_layout()
plt.show()
[占位-未完成]scikit-learn一般实例之十二:用于RBF核的显式特征映射逼近的更多相关文章
- C++面向对象类的实例题目十二
题目描述: 写一个程序计算正方体.球体和圆柱体的表面积和体积 程序代码: #include<iostream> #define PAI 3.1415 using namespace std ...
- 数据可视化实例(十二): 发散型条形图 (matplotlib,pandas)
https://datawhalechina.github.io/pms50/#/chapter10/chapter10 如果您想根据单个指标查看项目的变化情况,并可视化此差异的顺序和数量,那么散型条 ...
- Java开发笔记(七十二)Java8新增的流式处理
通过前面几篇文章的学习,大家应能掌握几种容器类型的常见用法,对于简单的增删改和遍历操作,各容器实例都提供了相应的处理方法,对于实际开发中频繁使用的清单List,还能利用Arrays工具的asList方 ...
- 框架源码系列十二:Mybatis源码之手写Mybatis
一.需求分析 1.Mybatis是什么? 一个半自动化的orm框架(Object Relation Mapping). 2.Mybatis完成什么工作? 在面向对象编程中,我们操作的都是对象,Myba ...
- [占位-未完成]scikit-learn一般实例之十:核岭回归和SVR的比较
[占位-未完成]scikit-learn一般实例之十:核岭回归和SVR的比较
- [占位-未完成]scikit-learn一般实例之十一:异构数据源的特征联合
[占位-未完成]scikit-learn一般实例之十一:异构数据源的特征联合 Datasets can often contain components of that require differe ...
- scikit learn 模块 调参 pipeline+girdsearch 数据举例:文档分类 (python代码)
scikit learn 模块 调参 pipeline+girdsearch 数据举例:文档分类数据集 fetch_20newsgroups #-*- coding: UTF-8 -*- import ...
- Scikit Learn: 在python中机器学习
转自:http://my.oschina.net/u/175377/blog/84420#OSC_h2_23 Scikit Learn: 在python中机器学习 Warning 警告:有些没能理解的 ...
- 《Android群英传》读书笔记 (5) 第十一章 搭建云端服务器 + 第十二章 Android 5.X新特性详解 + 第十三章 Android实例提高
第十一章 搭建云端服务器 该章主要介绍了移动后端服务的概念以及Bmob的使用,比较简单,所以略过不总结. 第十三章 Android实例提高 该章主要介绍了拼图游戏和2048的小项目实例,主要是代码,所 ...
随机推荐
- 【WCF】错误协定声明
在上一篇烂文中,老周给大伙伴们介绍了 IErrorHandler 接口的使用,今天,老周补充一个错误处理的知识点——错误协定. 错误协定与IErrorHandler接口不同,大伙伴们应该记得,上回我们 ...
- Ubuntu 16.10 开启PHP错误提示
两个步骤: 修改php.ini配置文件中的error_reporting 和 display_errors两地方内容: sudo vim /etc/php/7.0/apache2/php.ini er ...
- 利用XAG在RAC环境下实现GoldenGate自动Failover
概述 在RAC环境下配置OGG,要想实现RAC节点故障时,OGG能自动的failover到正常节点,要保证两点: 1. OGG的checkpoint,trail,BR文件放置在共享的集群文件系统上,R ...
- OpenCV模板匹配算法详解
1 理论介绍 模板匹配是在一幅图像中寻找一个特定目标的方法之一,这种方法的原理非常简单,遍历图像中的每一个可能的位置,比较各处与模板是否“相似”,当相似度足够高时,就认为找到了我们的目标.OpenCV ...
- PHP类和对象之重载
PHP中的重载指的是动态的创建属性与方法,是通过魔术方法来实现的.属性的重载通过__set,__get,__isset,__unset来分别实现对不存在属性的赋值.读取.判断属性是否设置.销毁属性. ...
- 脑洞大开之采用HTML5+SignalR2.0(.Net)实现原生Web视频
目录 对SignalR不了解的人可以直接移步下面的目录 SignalR系列目录 前言 - -,我又来了,今天废话不多说,我们直接来实现Web视频聊天. 采用的技术如下: HTML5 WebRTC Si ...
- BPM配置故事之案例12-触发另外流程
还记得阿海么,对就是之前的那个采购员,他又有了些意见. 阿海:小明,你看现在的流程让大家都这么方便,能不能帮个忙让我也轻松点啊-- 小明:--你有什么麻烦,现在不是已经各个部门自己提交申请了嘛? 阿海 ...
- AngularJS 系列 学习笔记 目录篇
目录: AngularJS 系列 01 - HelloWorld和数据绑定 AngularJS 系列 02 - 模块 (持续更新)
- Linux基础介绍【第三篇】
更改SSH服务端远程登录的配置 windows服务端的默认远程管理端口是3389,管理员用户是administrator,普通用户是guest.Linux的管理用户是root,普通用户默认有很多个,远 ...
- Web API 强势入门指南
Web API是一个比较宽泛的概念.这里我们提到Web API特指ASP.NET Web API. 这篇文章中我们主要介绍Web API的主要功能以及与其他同类型框架的对比,最后通过一些相对复杂的实例 ...
