Plotting trees from Random Forest models with ggraph
Today, I want to show how I use Thomas Lin Pederson’s awesome ggraph package to plot decision trees from Random Forest models.
I am very much a visual person, so I try to plot as much of my results as possible because it helps me get a better feel for what is going on with my data.
A nice aspect of using tree-based machine learning, like Random Forest models, is that that they are more easily interpreted than e.g. neural networks as they are based on decision trees. So, when I am using such models, I like to plot final decision trees (if they aren’t too large) to get a sense of which decisions are underlying my predictions.
There are a few very convient ways to plot the outcome if you are using the randomForest package but I like to have as much control as possible about the layout, colors, labels, etc. And because I didn’t find a solution I liked for caret models, I developed the following little function (below you may find information about how I built the model):
As input, it takes part of the output from model_rf <- caret::train(... "rf" ...), that gives the trees of the final model: model_rf$finalModel$forest. From these trees, you can specify which one to plot by index.
library(dplyr)
library(ggraph)
library(igraph)
tree_func <- function(final_model,
tree_num) {
# get tree by index
tree <- randomForest::getTree(final_model,
k = tree_num,
labelVar = TRUE) %>%
tibble::rownames_to_column() %>%
# make leaf split points to NA, so the 0s won't get plotted
mutate(`split point` = ifelse(is.na(prediction), `split point`, NA))
# prepare data frame for graph
graph_frame <- data.frame(from = rep(tree$rowname, 2),
to = c(tree$`left daughter`, tree$`right daughter`))
# convert to graph and delete the last node that we don't want to plot
graph <- graph_from_data_frame(graph_frame) %>%
delete_vertices("0")
# set node labels
V(graph)$node_label <- gsub("_", " ", as.character(tree$`split var`))
V(graph)$leaf_label <- as.character(tree$prediction)
V(graph)$split <- as.character(round(tree$`split point`, digits = 2))
# plot
plot <- ggraph(graph, 'dendrogram') +
theme_bw() +
geom_edge_link() +
geom_node_point() +
geom_node_text(aes(label = node_label), na.rm = TRUE, repel = TRUE) +
geom_node_label(aes(label = split), vjust = 2.5, na.rm = TRUE, fill = "white") +
geom_node_label(aes(label = leaf_label, fill = leaf_label), na.rm = TRUE,
repel = TRUE, colour = "white", fontface = "bold", show.legend = FALSE) +
theme(panel.grid.minor = element_blank(),
panel.grid.major = element_blank(),
panel.background = element_blank(),
plot.background = element_rect(fill = "white"),
panel.border = element_blank(),
axis.line = element_blank(),
axis.text.x = element_blank(),
axis.text.y = element_blank(),
axis.ticks = element_blank(),
axis.title.x = element_blank(),
axis.title.y = element_blank(),
plot.title = element_text(size = 18))
print(plot)
}We can now plot, e.g. the tree with the smalles number of nodes:
tree_num <- which(model_rf$finalModel$forest$ndbigtree == min(model_rf$finalModel$forest$ndbigtree))
tree_func(final_model = model_rf$finalModel, tree_num)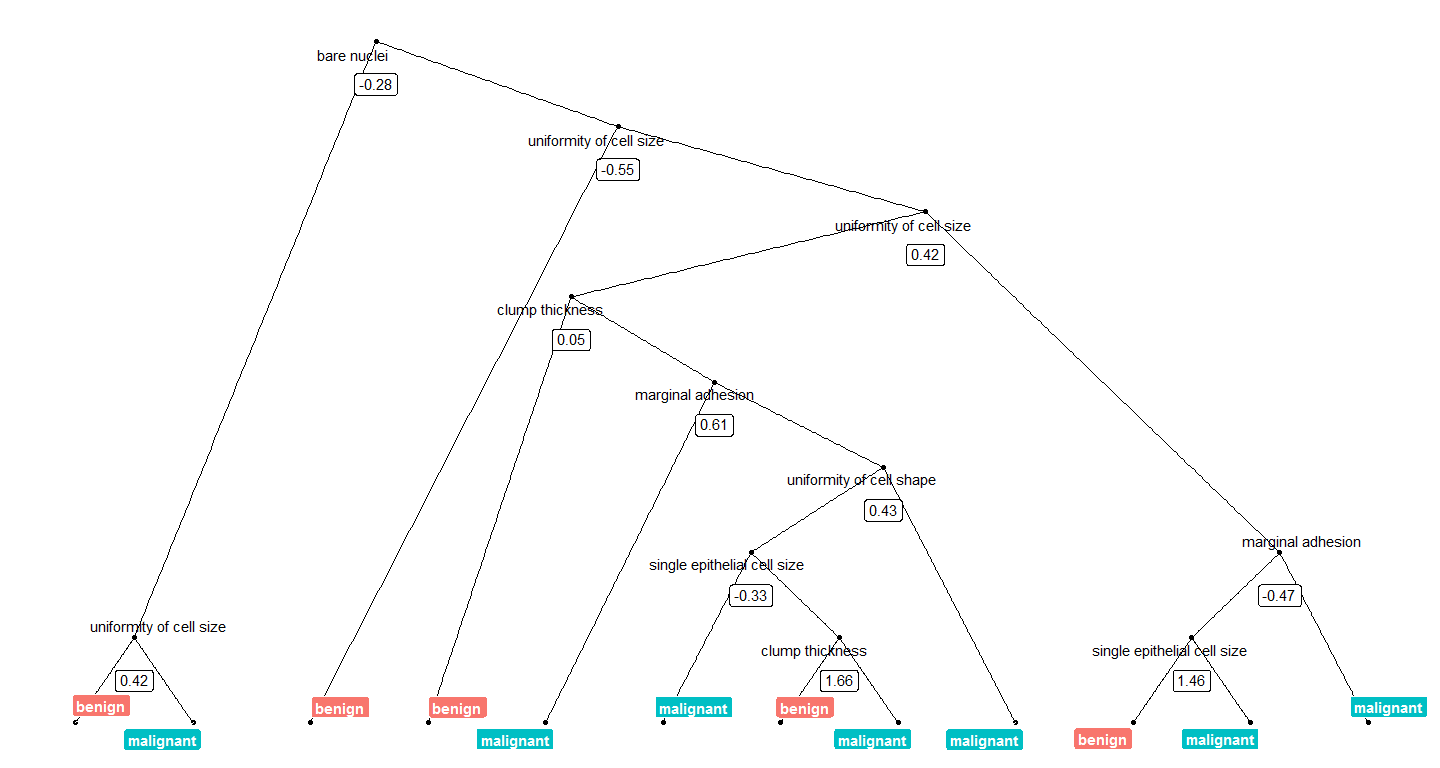
Or we can plot the tree with the biggest number of nodes:
tree_num <- which(model_rf$finalModel$forest$ndbigtree == max(model_rf$finalModel$forest$ndbigtree))
tree_func(final_model = model_rf$finalModel, tree_num)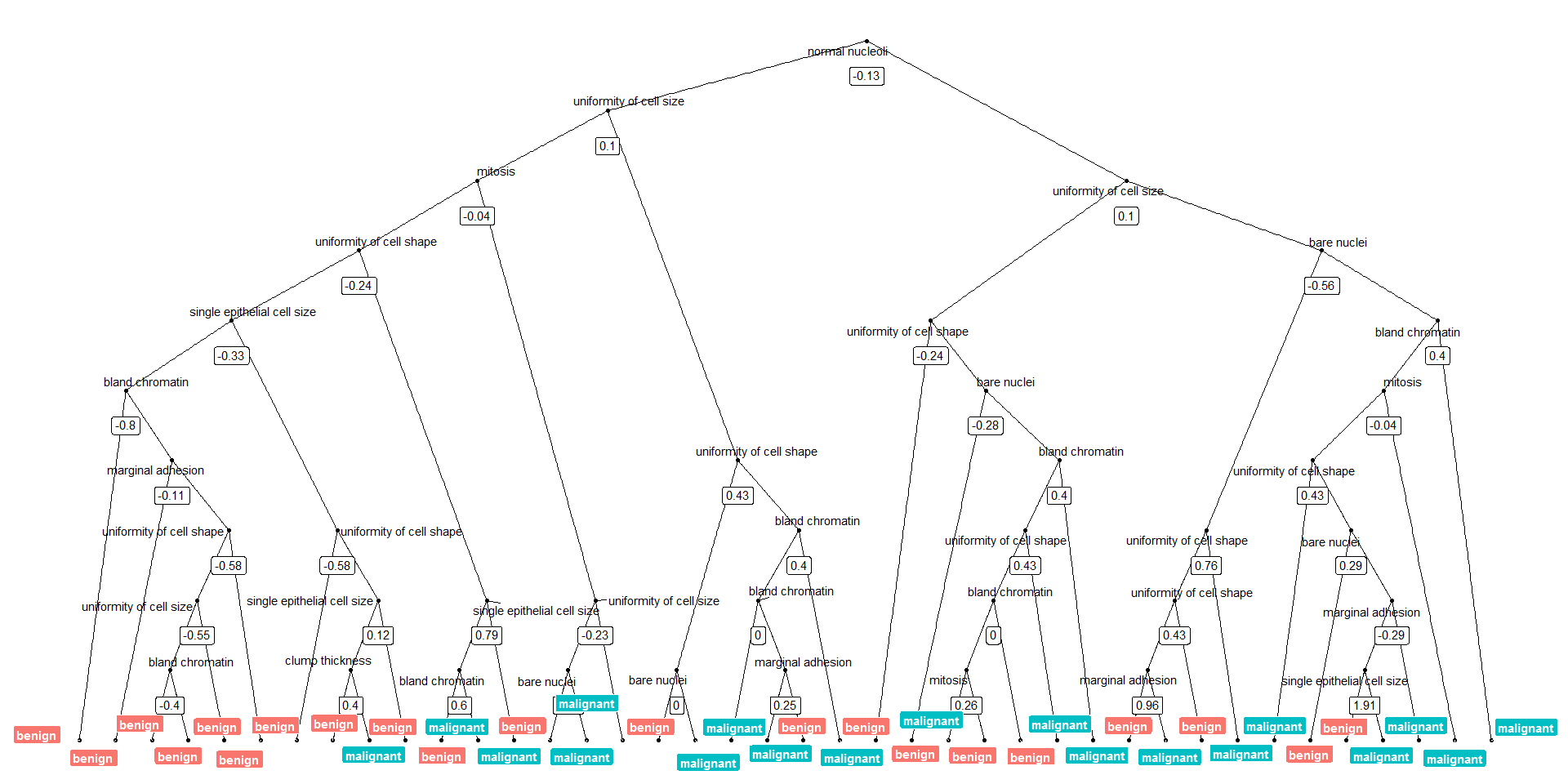
Preparing the data and modeling
The data set I am using in these example analyses, is the Breast Cancer Wisconsin (Diagnostic) Dataset. The data was downloaded from the UC Irvine Machine Learning Repository.
The first data set looks at the predictor classes:
- malignant or
- benign breast mass.
The features characterize cell nucleus properties and were generated from image analysis of fine needle aspirates (FNA) of breast masses:
- Sample ID (code number)
- Clump thickness
- Uniformity of cell size
- Uniformity of cell shape
- Marginal adhesion
- Single epithelial cell size
- Number of bare nuclei
- Bland chromatin
- Number of normal nuclei
- Mitosis
- Classes, i.e. diagnosis
bc_data <- read.table("datasets/breast-cancer-wisconsin.data.txt", header = FALSE, sep = ",")
colnames(bc_data) <- c("sample_code_number",
"clump_thickness",
"uniformity_of_cell_size",
"uniformity_of_cell_shape",
"marginal_adhesion",
"single_epithelial_cell_size",
"bare_nuclei",
"bland_chromatin",
"normal_nucleoli",
"mitosis",
"classes")
bc_data$classes <- ifelse(bc_data$classes == "2", "benign",
ifelse(bc_data$classes == "4", "malignant", NA))
bc_data[bc_data == "?"] <- NA
# impute missing data
library(mice)
bc_data[,2:10] <- apply(bc_data[, 2:10], 2, function(x) as.numeric(as.character(x)))
dataset_impute <- mice(bc_data[, 2:10], print = FALSE)
bc_data <- cbind(bc_data[, 11, drop = FALSE], mice::complete(dataset_impute, 1))
bc_data$classes <- as.factor(bc_data$classes)
# how many benign and malignant cases are there?
summary(bc_data$classes)
# separate into training and test data
library(caret)
set.seed(42)
index <- createDataPartition(bc_data$classes, p = 0.7, list = FALSE)
train_data <- bc_data[index, ]
test_data <- bc_data[-index, ]
# run model
set.seed(42)
model_rf <- caret::train(classes ~ .,
data = train_data,
method = "rf",
preProcess = c("scale", "center"),
trControl = trainControl(method = "repeatedcv",
number = 10,
repeats = 10,
savePredictions = TRUE,
verboseIter = FALSE))If you are interested in more machine learning posts, check out the category listing for machine_learning on my blog.
sessionInfo()## R version 3.3.3 (2017-03-06)
## Platform: x86_64-w64-mingw32/x64 (64-bit)
## Running under: Windows 7 x64 (build 7601) Service Pack 1
##
## locale:
## [1] LC_COLLATE=English_United States.1252
## [2] LC_CTYPE=English_United States.1252
## [3] LC_MONETARY=English_United States.1252
## [4] LC_NUMERIC=C
## [5] LC_TIME=English_United States.1252
##
## attached base packages:
## [1] stats graphics grDevices utils datasets methods base
##
## other attached packages:
## [1] igraph_1.0.1 ggraph_1.0.0 ggplot2_2.2.1.9000
## [4] dplyr_0.5.0
##
## loaded via a namespace (and not attached):
## [1] Rcpp_0.12.9 nloptr_1.0.4 plyr_1.8.4
## [4] viridis_0.3.4 iterators_1.0.8 tools_3.3.3
## [7] digest_0.6.12 lme4_1.1-12 evaluate_0.10
## [10] tibble_1.2 gtable_0.2.0 nlme_3.1-131
## [13] lattice_0.20-34 mgcv_1.8-17 Matrix_1.2-8
## [16] foreach_1.4.3 DBI_0.6 ggrepel_0.6.5
## [19] yaml_2.1.14 parallel_3.3.3 SparseM_1.76
## [22] gridExtra_2.2.1 stringr_1.2.0 knitr_1.15.1
## [25] MatrixModels_0.4-1 stats4_3.3.3 rprojroot_1.2
## [28] grid_3.3.3 caret_6.0-73 nnet_7.3-12
## [31] R6_2.2.0 rmarkdown_1.3 minqa_1.2.4
## [34] udunits2_0.13 tweenr_0.1.5 deldir_0.1-12
## [37] reshape2_1.4.2 car_2.1-4 magrittr_1.5
## [40] units_0.4-2 backports_1.0.5 scales_0.4.1
## [43] codetools_0.2-15 ModelMetrics_1.1.0 htmltools_0.3.5
## [46] MASS_7.3-45 splines_3.3.3 randomForest_4.6-12
## [49] assertthat_0.1 pbkrtest_0.4-6 ggforce_0.1.1
## [52] colorspace_1.3-2 labeling_0.3 quantreg_5.29
## [55] stringi_1.1.2 lazyeval_0.2.0 munsell_0.4.3转自:https://shiring.github.io/machine_learning/2017/03/16/rf_plot_ggraph
Plotting trees from Random Forest models with ggraph的更多相关文章
- 机器学习算法 --- Pruning (decision trees) & Random Forest Algorithm
一.Table for Content 在之前的文章中我们介绍了Decision Trees Agorithms,然而这个学习算法有一个很大的弊端,就是很容易出现Overfitting,为了解决此问题 ...
- Random Forest And Extra Trees
随机森林 我们对使用决策树随机取样的集成学习有个形象的名字–随机森林. scikit-learn 中封装的随机森林,在决策树的节点划分上,在随机的特征子集上寻找最优划分特征. import numpy ...
- Random Forest Classification of Mushrooms
There is a plethora of classification algorithms available to people who have a bit of coding experi ...
- 机器学习方法(六):随机森林Random Forest,bagging
欢迎转载,转载请注明:本文出自Bin的专栏blog.csdn.net/xbinworld. 技术交流QQ群:433250724,欢迎对算法.技术感兴趣的同学加入. 前面机器学习方法(四)决策树讲了经典 ...
- [Machine Learning & Algorithm] 随机森林(Random Forest)
1 什么是随机森林? 作为新兴起的.高度灵活的一种机器学习算法,随机森林(Random Forest,简称RF)拥有广泛的应用前景,从市场营销到医疗保健保险,既可以用来做市场营销模拟的建模,统计客户来 ...
- paper 85:机器统计学习方法——CART, Bagging, Random Forest, Boosting
本文从统计学角度讲解了CART(Classification And Regression Tree), Bagging(bootstrap aggregation), Random Forest B ...
- 多分类问题中,实现不同分类区域颜色填充的MATLAB代码(demo:Random Forest)
之前建立了一个SVM-based Ordinal regression模型,一种特殊的多分类模型,就想通过可视化的方式展示模型分类的效果,对各个分类区域用不同颜色表示.可是,也看了很多代码,但基本都是 ...
- 统计学习方法——CART, Bagging, Random Forest, Boosting
本文从统计学角度讲解了CART(Classification And Regression Tree), Bagging(bootstrap aggregation), Random Forest B ...
- sklearn_随机森林random forest原理_乳腺癌分类器建模(推荐AAA)
sklearn实战-乳腺癌细胞数据挖掘(博主亲自录制视频) https://study.163.com/course/introduction.htm?courseId=1005269003& ...
随机推荐
- Android中实现定时器的四种方式
第一种方式利用Timer和TimerTask 1.继承关系 java.util.Timer 基本方法 schedule 例如: timer.schedule(task, delay,period); ...
- 对InvokeRequired的理解
if (listBox1.InvokeRequired) //当有新工作进程访问控件时InvokeRequired为True ...
- AFNetworking 内部详解
AFNetworking 是一个适用于IOS 和 Mac OSX 两个平台的网络库,他是在Foundation URL Loading System 基础上进行的一套封装 ,并提供了丰富的API接口 ...
- 【算法】字符串匹配之Z算法
求文本与单模式串匹配,通常会使用KMP算法.后来接触到了Z算法,感觉Z算法也相当精妙.在以前的博文中也有过用Z算法来解决字符串匹配的题目. 下面介绍一下Z算法. 先一句话讲清楚Z算法能求什么东西. 输 ...
- Linux防火墙配置—SNAT1
1.实验目标 以实验"防火墙配置-访问外网WEB"为基础,在WEB服务器上安装Wireshark,设置Wireshark的过滤条件为捕获HTTP报文,在Wireshark中开启捕获 ...
- iOS 相册和网络图片的存取
iOS 相册和网络图片的存取 保存 UIImage 到相册 UIKit UIKit 中一个古老的方法,Objective-C 的形式 void UIImageWriteToSavedPhotosAlb ...
- DirectFB 之 实例图像不断右移
/********************************************** * Author: younger.liucn@gmail.com * File name: imgro ...
- MVC 5 + EF6 完整教程16 -- 控制器详解
Controller作为持久层和展现层的桥梁, 封装了应用程序的逻辑,是MVC中的核心组件之一. 本篇文章我们就来谈谈 Controller, 主要讨论两个方面: Controller运行机制简介 C ...
- java线程(二)
线程范围变量 我们知道线程在cpu上的使用权并不是长时间的,因为计算机的cpu只有一个,而在计算上运行的进程有很多,线程就更不用说了,所以cpu只能通过调度来上多个线程轮流占用cpu资源运行,且为了保 ...
- myeclipse/eclipse 配置SSM框架错误之一解决方法
报错如下: 1. [org.springframework.web.context.ContextLoader] - Root WebApplicationContext: initializatio ...
