dp-(LCS 基因匹配)
| Time Limit: 1000MS | Memory Limit: 10000K | |
| Total Submissions: 19885 | Accepted: 11100 |
Description
A human gene can be identified through a series of
time-consuming biological experiments, often with the help of computer
programs. Once a sequence of a gene is obtained, the next job is to
determine its function.
One of the methods for biologists to use in determining the function
of a new gene sequence that they have just identified is to search a
database with the new gene as a query. The database to be searched
stores many gene sequences and their functions – many researchers have
been submitting their genes and functions to the database and the
database is freely accessible through the Internet.
A database search will return a list of gene sequences from the database that are similar to the query gene.
Biologists assume that sequence similarity often implies
functional similarity. So, the function of the new gene might be
one of the functions that the genes from the list have. To exactly
determine which one is the right one another series of biological
experiments will be needed.
Your job is to make a program that compares two genes and determines
their similarity as explained below. Your program may be used as a part
of the database search if you can provide an efficient one.
Given two genes AGTGATG and GTTAG, how similar are they? One of the methods to measure the similarity
of two genes is called alignment. In an alignment, spaces are inserted, if necessary, in appropriate positions of
the genes to make them equally long and score the resulting genes according to a scoring matrix.
For example, one space is inserted into AGTGATG to result in
AGTGAT-G, and three spaces are inserted into GTTAG to result in
–GT--TAG. A space is denoted by a minus sign (-). The two genes
are now of equal
length. These two strings are aligned:
AGTGAT-G
-GT--TAG
In this alignment, there are four matches, namely, G in the second
position, T in the third, T in the sixth, and G in the eighth. Each
pair of aligned characters is assigned a score according to the
following scoring matrix.
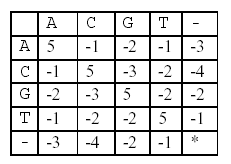
denotes that a space-space match is not allowed. The score
of the alignment above is (-3)+5+5+(-2)+(-3)+5+(-3)+5=9.
Of course, many other alignments are possible. One is shown below (a
different number of spaces are inserted into different positions):
AGTGATG
-GTTA-G
This alignment gives a score of (-3)+5+5+(-2)+5+(-1) +5=14. So,
this one is better than the previous one. As a matter of fact, this one
is optimal since no other alignment can have a higher score. So, it is
said that the
similarity of the two genes is 14.
Input
input consists of T test cases. The number of test cases ) (T is
given in the first line of the input file. Each test case consists of
two lines: each line contains an integer, the length of a gene, followed
by a gene sequence. The length of each gene sequence is at least one
and does not exceed 100.
Output
Sample Input
2
7 AGTGATG
5 GTTAG
7 AGCTATT
9 AGCTTTAAA
Sample Output
14
21 题目大意 : 给出一个基因匹配表格 , 里面有一些数值 。
设 dp[i][j] 为 s1 取第 i 个字符, s2 取第 j 个字符的最大值,决定dp[i][j] 最优的情况有 三种, 类似于最长公共子序列的三种情况。
1 . s1 取第 i 个字母, s2 取 '-' , temp1 = dp[i-1][j] + score[a[i]]['-'];
2 . s2 取第 j 个字母, s1 取 '-' , temp2 = dp[i][j-1] + score['-'][b[j]];
3 . s1 取第 i 个字母 , s2 取第 j 个字母 , temp3 = dp[i-1][j-1] + score[a[i]][b[j]]; 则 dp[i][j] = max ( temp1, temp2, temp3 ); 还有初始化问题 : dp[0][0] = 0;
dp[i][0] = dp[i-1][0] + score[a[i]]['-'];
dp[0][j] = dp[0][j-1] + score['-'][b[j]]; 代码示例 :
/*
* Author: ry
* Created Time: 2017/9/3 8:00:06
* File Name: 1.cpp
*/
#include <iostream>
#include <cstdio>
#include <cstdlib>
#include <cstring>
#include <cmath>
#include <algorithm>
#include <string>
#include <vector>
#include <stack>
#include <queue>
#include <set>
#include <time.h>
using namespace std;
const int mm = 1e6+5;
#define ll long long ll t_cnt;
void t_st(){t_cnt=clock();}
void t_ot(){printf("you spent : %lldms\n", clock()-t_cnt);}
//开始t_st();
//结束t_ot(); int dp[150][150];
int score['T'+1]['T'+1]; void intial () {
score['A']['A'] = 5;
score['C']['C'] = 5;
score['G']['G'] = 5;
score['T']['T'] = 5;
score['A']['C'] = score['C']['A'] = -1;
score['A']['G'] = score['G']['A'] = -2;
score['A']['T'] = score['T']['A'] = -1;
score['A']['-'] = score['-']['A'] = -3;
score['C']['G'] = score['G']['C'] = -3;
score['C']['T'] = score['T']['C'] = -2;
score['C']['-'] = score['-']['C'] = -4;
score['G']['T'] = score['T']['G'] = -2;
score['G']['-'] = score['-']['G'] = -2;
score['T']['-'] = score['-']['T'] = -1;
} int MAX (int x, int y, int z) {
int k = (x>y?x:y);
return z>k?z:k;
} int main() {
int t;
int len1, len2;
char a[105], b[105]; intial();
cin >> t;
getchar();
while ( t-- ){
memset (dp, 0, sizeof(dp));
scanf("%d %s", &len1, a);
scanf("%d %s", &len2, b); for (int i = len1; i > 0; i--)
a[i] = a[i-1];
for (int i = len2; i > 0; i--)
b[i] = b[i-1]; dp[0][0] = 0;
for (int i = 1; i <= len1; i++)
dp[i][0] = dp[i-1][0] + score[a[i]]['-'];
for (int j = 1; j <= len2; j++)
dp[0][j] = dp[0][j-1] + score['-'][b[j]]; for (int i = 1; i <= len1; i++)
for (int j = 1; j <= len2; j++){
int temp1 = dp[i-1][j] + score[a[i]]['-'];
int temp2 = dp[i][j-1] + score['-'][b[j]];
int temp3 = dp[i-1][j-1] + score[a[i]][b[j]];
dp[i][j] = MAX (temp1, temp2, temp3);
} printf ("%d\n", dp[len1][len2]);
} return 0;
}
dp-(LCS 基因匹配)的更多相关文章
- 【线型DP】【LCS】洛谷P4303 [AHOI2006]基因匹配
P4303 [AHOI2006]基因匹配 标签(空格分隔): 考试题 nt题 LCS优化 [题目] 卡卡昨天晚上做梦梦见他和可可来到了另外一个星球,这个星球上生物的DNA序列由无数种碱基排列而成(地球 ...
- BZOJ 1264: [AHOI2006]基因匹配Match( LCS )
序列最大长度2w * 5 = 10w, O(n²)的LCS会T.. LCS 只有当a[i] == b[j]时, 才能更新答案, 我们可以记录n个数在第一个序列中出现的5个位置, 然后从左往右扫第二个序 ...
- bzoj 1264 [AHOI2006]基因匹配Match(DP+树状数组)
1264: [AHOI2006]基因匹配Match Time Limit: 10 Sec Memory Limit: 162 MBSubmit: 793 Solved: 503[Submit][S ...
- bzoj1264 [AHOI2006]基因匹配Match 树状数组+lcs
1264: [AHOI2006]基因匹配Match Time Limit: 10 Sec Memory Limit: 162 MBSubmit: 1255 Solved: 835[Submit][ ...
- BZOJ 1264: [AHOI2006]基因匹配Match 树状数组+DP
1264: [AHOI2006]基因匹配Match Description 基因匹配(match) 卡卡昨天晚上做梦梦见他和可可来到了另外一个星球,这个星球上生物的DNA序列由无数种碱基排列而成(地球 ...
- 【BZOJ1264】[AHOI2006]基因匹配Match DP+树状数组
[BZOJ1264][AHOI2006]基因匹配Match Description 基因匹配(match) 卡卡昨天晚上做梦梦见他和可可来到了另外一个星球,这个星球上生物的DNA序列由无数种碱基排列而 ...
- 基因匹配(bzoj 1264)
Description 基因匹配(match) 卡卡昨天晚上做梦梦见他和可可来到了另外一个星球,这个星球上生物的DNA序列由无数种碱基排列而成(地球上只有4种),而更奇怪的是,组成DNA序列的每一种碱 ...
- BZOJ1264: [AHOI2006]基因匹配Match
1264: [AHOI2006]基因匹配Match Time Limit: 10 Sec Memory Limit: 162 MBSubmit: 541 Solved: 347[Submit][S ...
- [AHOI2006]基因匹配
题目描述 卡卡昨天晚上做梦梦见他和可可来到了另外一个星球,这个星球上生物的DNA序列由无数种碱基排列而成(地球上只有4种),而更奇怪的是,组成DNA序列的每一种碱基在该序列中正好出现5次!这样如果一个 ...
- bzoj 1264: [AHOI2006]基因匹配Match
1264: [AHOI2006]基因匹配Match Description 基因匹配(match) 卡卡昨天晚上做梦梦见他和可可来到了另外一个星球,这个星球上生物的DNA序列由无数种碱基排列而成(地球 ...
随机推荐
- 配置gitignore后使其生效命令
改动过.gitignore文件之后,在repo的根目录下运行: git rm -r --cached . git add . 之后可以进行提交: git commit -m "fixed u ...
- C# 命令行如何静默调用 del 删除文件
如果在 C# 命令行调用 del 删除文件,很多时候会提示是否需要删除,本文告诉大家如何调用命令行的时候静默删除 在C# 命令行 调用 del 删除文件的时候,会提示是否删除,通过在命令行加上 \Q ...
- Filter、Intercepter、AOP的区别
在使用Spring MVC开发RESTful API的时候,我们经常会使用Java的拦截机制来处理请求,Filter是Java本身自带拦过滤器,Interceptor则是Spring自带的拦截器,而A ...
- linux 安装一个中断处理
如果你想实际地"看到"产生的中断, 向硬件设备写不足够; 一个软件处理必须在系统中配 置. 如果 Linux 内核还没有被告知来期待你的中断, 它简单地确认并忽略它. 中断线是一个 ...
- 大数据基石——Hadoop与MapReduce
本文始发于个人公众号:TechFlow 近两年AI成了最火热领域的代名词,各大高校纷纷推出了人工智能专业.但其实,人工智能也好,还是前两年的深度学习或者是机器学习也罢,都离不开底层的数据支持.对于动辄 ...
- 0018 CSS注释(简单)
CSS注释规则: /* 需要注释的内容 */ 进行注释的,即在需要注释的内容前使用 "/*" 标记开始注释,在内容的结尾使用 "*/"结束. 例如: p { / ...
- Linux 批量安装依赖
1.依赖检测失败,xxx被xxxx需要. 当我安装rpm 的时候,出现依赖检测失败. 我们可以到http://rpmfind.net/linux/rpm2html/search.php 这个网站上去搜 ...
- DOCKER学习_010:Docker的文件系统以及制作镜像
一 文件系统简介 1.1 Linux文件系统 LInux空间组成分为内核空间和用户空间(使用rootfs) linux文件系统由 bootes和 rootfs组成, bootes主要包含boot1 o ...
- alpha week 2/2 Scrum立会报告+燃尽图 06
此作业要求参见:https://edu.cnblogs.com/campus/nenu/2019fall/homework/9803 小组名称:“组长”组 组长:杨天宇 组员:魏新,罗杨美慧,王歆瑶, ...
- vc调用mysql数据库操作例子
这里归纳了C API可使用的函数 函数 描述 mysql_affected_rows() 返回上次UPDATE.DELETE或INSERT查询更改/删除/插入的行数. mysql_autocommit ...
