plot a critical difference diagram , MATLAB code
plot a critical difference diagram , MATLAB code
建立criticaldifference函数
function cd = criticaldifference(s,labels,alpha)
%
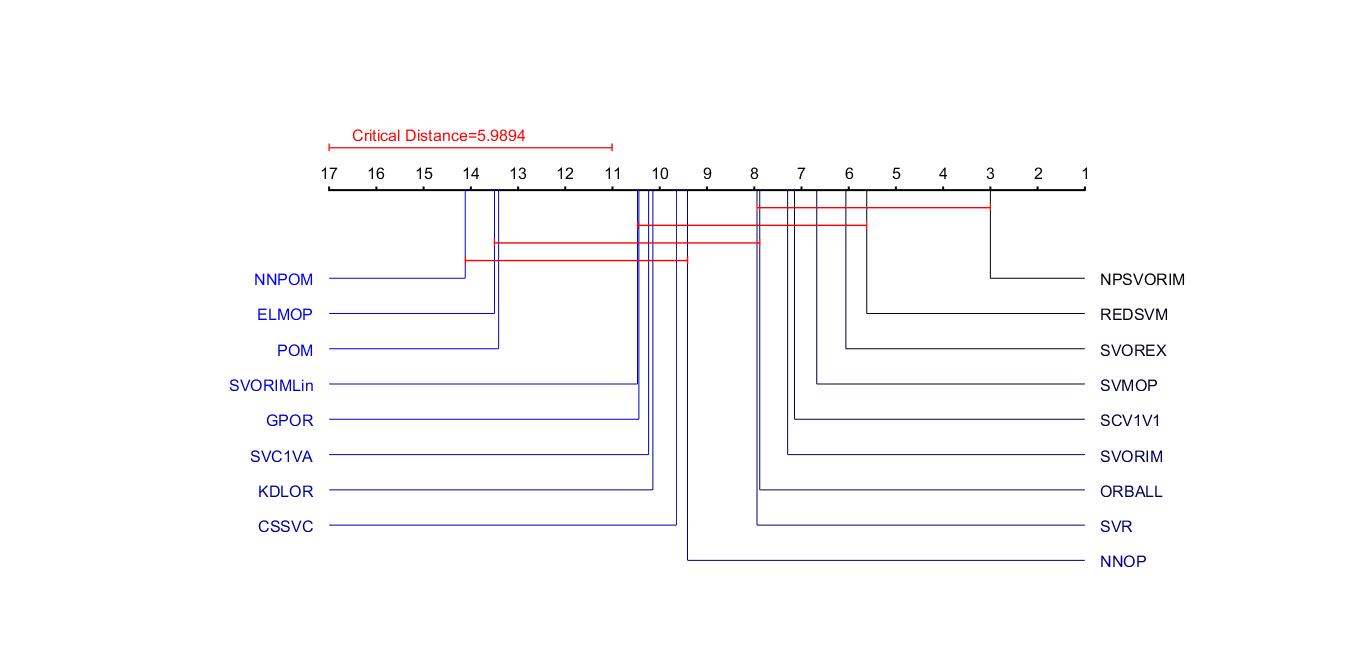
% CRITICALDIFFERNCE - plot a critical difference diagram
%
% CRITICALDIFFERENCE(S,LABELS) produces a critical difference diagram [1]
% displaying the statistical significance (or otherwise) of a matrix of
% scores, S, achieved by a set of machine learning algorithms. Here
% LABELS is a cell array of strings giving the name of each algorithm.
%
% References
%
% [1] Demsar, J., "Statistical comparisons of classifiers over multiple
% datasets", Journal of Machine Learning Research, vol. 7, pp. 1-30,
% 2006.
% %
% File : criticaldifference.m
%
% Date : Monday 14th April 2008
%
% Author : Gavin C. Cawley
%
% Description : Sparse multinomial logistic regression using a Laplace prior.
%
% History : 14/04/2008 - v1.00
%
% Copyright : (c) Dr Gavin C. Cawley, April 2008.
%
% This program is free software; you can redistribute it and/or modify
% it under the terms of the GNU General Public License as published by
% the Free Software Foundation; either version 2 of the License, or
% (at your option) any later version.
%
% This program is distributed in the hope that it will be useful,
% but WITHOUT ANY WARRANTY; without even the implied warranty of
% MERCHANTABILITY or FITNESS FOR A PARTICULAR PURPOSE. See the
% GNU General Public License for more details.
%
% You should have received a copy of the GNU General Public License
% along with this program; if not, write to the Free Software
% Foundation, Inc., 59 Temple Place, Suite 330, Boston, MA 02111-1307 USA
% % Thanks to Gideon Dror for supplying the extended table of critical values. if nargin < 3
alpha = 0.1;
end % convert scores into ranks
[N,k] = size(s);
[S,r] = sort(s');
idx = k*repmat(0:N-1, k, 1)' + r';
R = repmat(1:k, N, 1);
S = S'; for i=1:N
for j=1:k
index = S(i,j) == S(i,:);
R(i,index) = mean(R(i,index));
end
end r(idx) = R;
r = r'; % compute critical difference
if alpha == 0.01
qalpha = [0.000 2.576 2.913 3.113 3.255 3.364 3.452 3.526 3.590 3.646 ...
3.696 3.741 3.781 3.818 3.853 3.884 3.914 3.941 3.967 3.992 ...
4.015 4.037 4.057 4.077 4.096 4.114 4.132 4.148 4.164 4.179 ...
4.194 4.208 4.222 4.236 4.249 4.261 4.273 4.285 4.296 4.307 ...
4.318 4.329 4.339 4.349 4.359 4.368 4.378 4.387 4.395 4.404 ...
4.412 4.420 4.428 4.435 4.442 4.449 4.456 ]; elseif alpha == 0.05
qalpha = [0.000 1.960 2.344 2.569 2.728 2.850 2.948 3.031 3.102 3.164 ...
3.219 3.268 3.313 3.354 3.391 3.426 3.458 3.489 3.517 3.544 ...
3.569 3.593 3.616 3.637 3.658 3.678 3.696 3.714 3.732 3.749 ...
3.765 3.780 3.795 3.810 3.824 3.837 3.850 3.863 3.876 3.888 ...
3.899 3.911 3.922 3.933 3.943 3.954 3.964 3.973 3.983 3.992 ...
4.001 4.009 4.017 4.025 4.032 4.040 4.046]; elseif alpha == 0.1
qalpha = [0.000 1.645 2.052 2.291 2.460 2.589 2.693 2.780 2.855 2.920 ...
2.978 3.030 3.077 3.120 3.159 3.196 3.230 3.261 3.291 3.319 ...
3.346 3.371 3.394 3.417 3.439 3.459 3.479 3.498 3.516 3.533 ...
3.550 3.567 3.582 3.597 3.612 3.626 3.640 3.653 3.666 3.679 ...
3.691 3.703 3.714 3.726 3.737 3.747 3.758 3.768 3.778 3.788 ...
3.797 3.806 3.814 3.823 3.831 3.838 3.846]; else
error('alpha must be 0.01, 0.05 or 0.1');
end cd = qalpha(k)*sqrt(k*(k+1)/(6*N)); figure(1);
clf
axis off
axis([-0.2 1.2 -20 140]);
axis xy
tics = repmat((0:(k-1))/(k-1), 3, 1);
line(tics(:), repmat([100, 101, 100], 1, k), 'LineWidth', 1.5, 'Color', 'k');
%tics = repmat(((0:(k-2))/(k-1)) + 0.5/(k-1), 3, 1);
%line(tics(:), repmat([100, 101, 100], 1, k-1), 'LineWidth', 1.5, 'Color', 'k');
line([0 0 0 cd/(k-1) cd/(k-1) cd/(k-1)], [113 111 112 112 111 113], 'LineWidth', 1, 'Color', 'r');
text(0.03, 116, ['Critical Distance=' num2str(cd)], 'FontSize', 12, 'HorizontalAlignment', 'left', 'Color', 'r'); for i=1:k
text((i-1)/(k-1), 105, num2str(k-i+1), 'FontSize', 12, 'HorizontalAlignment', 'center');
end % compute average ranks
r = mean(r);
[r,idx] = sort(r); % compute statistically similar cliques
clique = repmat(r,k,1) - repmat(r',1,k);
clique(clique<0) = realmax;
clique = clique < cd; for i=k:-1:2
if all(clique(i-1,clique(i,:))==clique(i,clique(i,:)))
clique(i,:) = 0;
end
end n = sum(clique,2);
clique = clique(n>1,:);
n = size(clique,1); %yanse={'b','g','y','m','r'};
b=linspace(0,1,k);
% labels displayed on the right
for i=1:ceil(k/2)
line([(k-r(i))/(k-1) (k-r(i))/(k-1) 1], [100 100-3*(n+1)-10*i 100-3*(n+1)-10*i], 'Color', [0 0 b(i)]);
%text(1.2, 100 - 5*(n+1)- 10*i + 2, num2str(r(i)), 'FontSize', 10, 'HorizontalAlignment', 'right');
text(1.02, 100 - 3*(n+1) - 10*i, labels{idx(i)}, 'FontSize', 12, 'VerticalAlignment', 'middle', 'HorizontalAlignment', 'left', 'Color', [0 0 b(i)]);
end % labels displayed on the left
for i=ceil(k/2)+1:k
line([(k-r(i))/(k-1) (k-r(i))/(k-1) 0], [100 100-3*(n+1)-10*(k-i+1) 100-3*(n+1)-10*(k-i+1)], 'Color', [0 0 b(i)]);
%text(-0.2, 100 - 5*(n+1) -10*(k-i+1)+2, num2str(r(i)), 'FontSize', 10, 'HorizontalAlignment', 'left');
text(-0.02, 100 - 3*(n+1) -10*(k-i+1), labels{idx(i)}, 'FontSize', 12, 'VerticalAlignment', 'middle', 'HorizontalAlignment', 'right', 'Color', [0 0 b(i)]);
end % group cliques of statistically similar classifiers
for i=1:size(clique,1)
R = r(clique(i,:));
%line([((k-min(R))/(k-1)) + 0.015 ((k - max(R))/(k-1)) - 0.015], [100-5*i 100-5*i], 'LineWidth', 1, 'Color', 'r');
%line([0 0 0 cd/(k-1) cd/(k-1) cd/(k-1)], [113 111 112 112 111 113], 'LineWidth', 1, 'Color', 'r');
line([((k-min(R))/(k-1)) ((k-min(R))/(k-1)) ((k-min(R))/(k-1)) ((k - max(R))/(k-1)) ((k - max(R))/(k-1)) ((k - max(R))/(k-1))], [100+1-5*i 100-1-5*i 100-5*i 100-5*i 100-1-5*i 100+1-5*i], 'LineWidth', 1, 'Color', 'r');
end
可执行m文件:
load Data
s=AccMatrix;
labels={'SCV1V1','SVC1VA','SVR','CSSVC','SVMOP','NNOP','ELMOP','POM',...
'NNPOM', 'SVOREX','SVORIM','SVORIMLin','KDLOR','GPOR','REDSVM','ORBALL' ,'NPSVORIM'};%方法的标签 alpha=0.05; %显著性水平0.1,0.05或0.01
cd = criticaldifference(s,labels,alpha)
AccMatrix=[
0.28 0.12 0.28 0.11 0.32 0.08 0.26 0.13 0.37 0.10 0.28 0.12 0.42 0.21 0.38 0.17 0.36 0.14 0.36 0.13 0.38 0.12 0.37 0.10 0.34 0.15 0.39 0.09 0.37 0.12 0.36 0.13 0.37 0.11
0.31 0.12 0.33 0.11 0.34 0.13 0.32 0.11 0.32 0.09 0.24 0.11 0.40 0.18 0.50 0.15 0.34 0.18 0.35 0.12 0.34 0.12 0.34 0.12 0.33 0.11 0.48 0.17 0.33 0.11 0.30 0.12 0.28 0.14
0.36 0.09 0.40 0.14 0.39 0.11 0.39 0.13 0.40 0.09 0.39 0.11 0.44 0.16 0.62 0.15 0.50 0.13 0.37 0.13 0.37 0.13 0.37 0.13 0.39 0.12 0.55 0.10 0.38 0.13 0.36 0.12 0.32 0.10
0.22 0.12 0.28 0.16 0.24 0.10 0.27 0.15 0.27 0.11 0.29 0.11 0.39 0.13 0.65 0.14 0.39 0.14 0.26 0.11 0.27 0.11 0.32 0.11 0.26 0.11 0.36 0.16 0.27 0.12 0.30 0.10 0.22 0.10
0.44 0.06 0.45 0.06 0.40 0.07 0.43 0.07 0.46 0.06 0.41 0.06 0.44 0.08 0.50 0.08 0.45 0.09 0.41 0.07 0.40 0.07 0.48 0.07 0.43 0.05 0.67 0.04 0.40 0.07 0.40 0.06 0.41 0.05
0.03 0.03 0.04 0.03 0.04 0.02 0.04 0.02 0.04 0.03 0.04 0.02 0.06 0.02 0.03 0.02 0.03 0.03 0.03 0.02 0.03 0.02 0.03 0.02 0.03 0.02 0.03 0.02 0.03 0.02 0.04 0.03 0.03 0.03
0.03 0.01 0.03 0.01 0.16 0.03 0.03 0.01 0.03 0.01 0.04 0.01 0.09 0.02 0.09 0.02 0.06 0.05 0.00 0.01 0.00 0.01 0.09 0.02 0.16 0.03 0.03 0.01 0.00 0.00 0.03 0.02 0.02 0.01
0.42 0.03 0.44 0.03 0.43 0.03 0.43 0.03 0.42 0.03 0.42 0.03 0.43 0.02 0.43 0.03 0.46 0.03 0.43 0.03 0.43 0.03 0.43 0.03 0.51 0.03 0.42 0.03 0.43 0.03 0.44 0.03 0.43 0.03
0.01 0.00 0.01 0.01 0.03 0.01 0.01 0.01 0.00 0.00 0.03 0.01 0.16 0.01 0.84 0.30 0.11 0.02 0.01 0.01 0.01 0.01 0.08 0.01 0.05 0.01 0.04 0.01 0.01 0.00 0.01 0.01 0.01 0.00
0.43 0.04 0.43 0.06 0.46 0.07 0.43 0.06 0.45 0.10 0.53 0.09 0.57 0.13 0.66 0.16 0.62 0.14 0.45 0.06 0.45 0.07 0.43 0.08 0.47 0.09 0.42 0.03 0.44 0.05 0.46 0.09 0.42 0.08
0.05 0.03 0.05 0.03 0.07 0.04 0.05 0.03 0.07 0.03 0.06 0.03 0.07 0.03 0.71 0.03 0.06 0.03 0.02 0.01 0.02 0.01 0.74 0.01 0.11 0.03 0.05 0.02 0.02 0.01 0.05 0.02 0.04 0.03
0.36 0.03 0.45 0.03 0.36 0.03 0.44 0.03 0.35 0.03 0.42 0.04 0.43 0.03 0.85 0.02 0.46 0.04 0.36 0.03 0.36 0.03 0.36 0.02 0.37 0.03 0.31 0.03 0.36 0.03 0.38 0.03 0.34 0.03
0.37 0.02 0.37 0.02 0.38 0.02 0.37 0.02 0.37 0.02 0.37 0.03 0.37 0.02 0.38 0.03 0.38 0.02 0.38 0.02 0.38 0.02 0.39 0.02 0.46 0.03 0.39 0.03 0.37 0.02 0.39 0.03 0.37 0.03
0.25 0.06 0.26 0.06 0.32 0.07 0.27 0.06 0.26 0.04 0.39 0.06 0.38 0.06 0.53 0.19 0.55 0.08 0.32 0.05 0.32 0.07 0.41 0.07 0.30 0.07 0.39 0.07 0.32 0.07 0.29 0.05 0.27 0.05
0.35 0.02 0.36 0.02 0.37 0.02 0.36 0.02 0.36 0.02 0.40 0.02 0.40 0.02 0.40 0.02 0.40 0.02 0.37 0.02 0.37 0.02 0.41 0.02 0.35 0.02 0.39 0.01 0.37 0.02 0.33 0.02 0.36 0.02
0.31 0.04 0.33 0.03 0.30 0.03 0.32 0.03 0.29 0.03 0.31 0.04 0.30 0.04 0.29 0.03 0.34 0.13 0.29 0.03 0.28 0.03 0.29 0.04 0.36 0.03 0.29 0.03 0.29 0.03 0.32 0.02 0.29 0.03
0.74 0.02 0.82 0.03 0.75 0.02 0.80 0.03 0.74 0.02 0.71 0.02 0.75 0.02 0.74 0.02 0.73 0.03 0.71 0.03 0.75 0.02 0.76 0.02 0.81 0.03 0.71 0.03 0.75 0.02 0.76 0.02 0.75 0.03 ];
plot a critical difference diagram , MATLAB code的更多相关文章
- Silence Removal and End Point Detection MATLAB Code
转载自:http://ganeshtiwaridotcomdotnp.blogspot.com/2011/08/silence-removal-and-end-point-detection.html ...
- Compute Mean Value of Train and Test Dataset of Caltech-256 dataset in matlab code
Compute Mean Value of Train and Test Dataset of Caltech-256 dataset in matlab code clc;imPath = '/ho ...
- Matlab Code for Visualize the Tracking Results of OTB100 dataset
Matlab Code for Visualize the Tracking Results of OTB100 dataset 2018-11-12 17:06:21 %把所有tracker的结果画 ...
- 支持向量机的smo算法(MATLAB code)
建立smo.m % function [alpha,bias] = smo(X, y, C, tol) function model = smo(X, y, C, tol) % SMO: SMO al ...
- MFCC matlab code
%function ccc=mfcc(x) %归一化mel滤波器组系数 filename=input('input filename:','s'); [x,fs,bits]=wavread(filen ...
- word linkage 选择合适的聚类个数matlab code
clear load fisheriris X = meas; m = size(X,2); % load machine % load census % % X = meas; % X=X(1:20 ...
- sequential minimal optimization,SMO for SVM, (MATLAB code)
function model = SMOforSVM(X, y, C ) %sequential minimal optimization,SMO tol = 0.001; maxIters = 30 ...
- MATLAB中矢量场图的绘制 (quiver/quiver3/dfield/pplane) Plot the vector field with MATLAB
1.quiver函数 一般用于绘制二维矢量场图,函数调用方法如下: quiver(x,y,u,v) 该函数展示了点(x,y)对应的的矢量(u,v).其中,x的长度要求等于u.v的列数,y的长度要求等于 ...
- 求平均排序MATLAB code
A0=R(:,1:2:end); for i=1:17 A1=A0(i,:); p=sort(unique(A1)); for j=1:length(p) Rank0(A1==p(j))=j; end ...
随机推荐
- [Effective Java]第十一章 序列化
声明:原创作品,转载时请注明文章来自SAP师太技术博客( 博/客/园www.cnblogs.com):www.cnblogs.com/jiangzhengjun,并以超链接形式标明文章原始出处,否则将 ...
- So easy Webservice 8.spring整合CXF 发布WS
1.添加jar包(cxf的jar包中包含了spring的jar包),添加spring配置文件 2.web.xml中配置CXFServlet,过滤WS服务的地址 <!-- 配置CXFServlet ...
- android测试参考,及CreateProcess failure, error问题解决
今天小伙伴问我问题,我给了这2个小命令,或许做android测试的同学可以用得着. 截图命令adb shell /system/bin/screencap -p /sdcard/screenshot. ...
- HDU1016 Prime Ring Problem(DFS回溯)
Prime Ring Problem Time Limit: 4000/2000 MS (Java/Others) Memory Limit: 65536/32768 K (Java/Other ...
- MyEclipse Servers视窗出现“Could not create the view: An unexpected exception was thrown”错误解决办法
打开所在的wordspace文件夹,在下面子文件夹 .metadata\.plugins\org.eclipse.core.runtime\.settings\com.genuitec.eclipse ...
- Data truncation: Truncated incorrect DOUBLE value 解决方案
1.情况限制 此处的错误解决方案只讨论: 在使用Mybatis时,传入数组且使用<foreach>标签时出现此种报错: 2.报错案例 mapper.java /** * @Descript ...
- SQL & PL/SQL 模块总结
SQL 1. 各种function 2. merge 3. connect by PL/SQL 1. pl/sql 寄出 2. 游标 3. procedure 4. function 5. packa ...
- Java 多线程 (转)
http://www.ibm.com/developerworks/cn/java/j-thread/index.html http://www.ibm.com/developerworks/cn/j ...
- Android批量图片加载经典系列——采用二级缓存、异步加载网络图片
一.问题描述 Android应用中经常涉及从网络中加载大量图片,为提升加载速度和效率,减少网络流量都会采用二级缓存和异步加载机制,所谓二级缓存就是通过先从内存中获取.再从文件中获取,最后才会访问网络. ...
- (转)Thread.setDaemon设置说明
本想搜下python多线程里的setDaemon,发现了这篇文章写得很不错:http://blog.csdn.net/m13666368773/article/details/7245570 Thre ...
