POJ1080(LCS变形)
Human Gene Functions
Time Limit: 2000/1000 MS (Java/Others) Memory Limit: 65536/32768 K (Java/Others)
Total Submission(s): 2799 Accepted Submission(s): 1587
A human gene can be identified through a series of time-consuming biological experiments, often with the help of computer programs. Once a sequence of a gene is obtained, the next job is to determine its function. One of the methods for biologists to use in determining the function of a new gene sequence that they have just identified is to search a database with the new gene as a query. The database to be searched stores many gene sequences and their functions – many researchers have been submitting their genes and functions to the database and the database is freely accessible through the Internet.
A database search will return a list of gene sequences from the database that are similar to the query gene. Biologists assume that sequence similarity often implies functional similarity. So, the function of the new gene might be one of the functions that the genes from the list have. To exactly determine which one is the right one another series of biological experiments will be needed.
Your job is to make a program that compares two genes and determines their similarity as explained below. Your program may be used as a part of the database search if you can provide an efficient one.
Given two genes AGTGATG and GTTAG, how similar are they? One of the methods to measure the similarity of two genes is called alignment. In an alignment, spaces are inserted, if necessary, in appropriate positions of the genes to make them equally long and score the resulting genes according to a scoring matrix.
For example, one space is inserted into AGTGATG to result in AGTGAT-G, and three spaces are inserted into GTTAG to result in –GT--TAG. A space is denoted by a minus sign (-). The two genes are now of equal length. These two strings are aligned:
AGTGAT-G
-GT--TAG
In this alignment, there are four matches, namely, G in the second position, T in the third, T in the sixth, and G in the eighth. Each pair of aligned characters is assigned a score according to the following scoring matrix.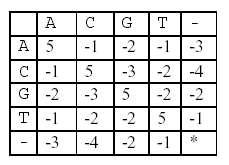
* denotes that a space-space match is not allowed. The score of the alignment above is (-3)+5+5+(-2)+(-3)+5+(-3)+5=9.
Of course, many other alignments are possible. One is shown below (a different number of spaces are inserted into different positions):
AGTGATG
-GTTA-G
This alignment gives a score of (-3)+5+5+(-2)+5+(-1) +5=14. So, this one is better than the previous one. As a matter of fact, this one is optimal since no other alignment can have a higher score. So, it is said that the similarity of the two genes is 14.
/*
ID: LinKArftc
PROG: 1080.cpp
LANG: C++
*/ #include <map>
#include <set>
#include <cmath>
#include <stack>
#include <queue>
#include <vector>
#include <cstdio>
#include <string>
#include <utility>
#include <cstdlib>
#include <cstring>
#include <iostream>
#include <algorithm>
using namespace std;
#define eps 1e-8
#define randin srand((unsigned int)time(NULL))
#define input freopen("input.txt","r",stdin)
#define debug(s) cout << "s = " << s << endl;
#define outstars cout << "*************" << endl;
const double PI = acos(-1.0);
const double e = exp(1.0);
const int inf = 0x3f3f3f3f;
const int INF = 0x7fffffff;
typedef long long ll; const int maxn = ;
int score[][] = {
{, -, -, -, -},
{-, , -, -, -},
{-, -, , -, -},
{-, -, -, , -},
{-, -, -, -, }
}; char ch_str1[maxn], ch_str2[maxn];
int str1[maxn], str2[maxn];
int dp[maxn][maxn]; int main() {
int T;
int len1, len2;
scanf("%d", &T);
while (T --) {
scanf("%d %s", &len1, ch_str1);
scanf("%d %s", &len2, ch_str2);
for (int i = ; i < len1; i ++) {
if (ch_str1[i] == 'A') str1[i] = ;
else if (ch_str1[i] == 'C') str1[i] = ;
else if (ch_str1[i] == 'G') str1[i] = ;
else if (ch_str1[i] == 'T') str1[i] = ;
else str1[i] = ;
}
for (int i = ; i < len2; i ++) {
if (ch_str2[i] == 'A') str2[i] = ;
else if (ch_str2[i] == 'C') str2[i] = ;
else if (ch_str2[i] == 'G') str2[i] = ;
else if (ch_str2[i] == 'T') str2[i] = ;
else str2[i] = ;
}
memset(dp, , sizeof(dp));
for (int i = ; i <= len1; i ++) dp[i][] = dp[i-][] + score[str1[i-]][];
for (int i = ; i <= len2; i ++) dp[][i] = dp[][i-] + score[][str2[i-]];
for (int i = ; i <= len1; i ++) {
for (int j = ; j <= len2; j ++) {
dp[i][j] = max(dp[i-][j-] + score[str1[i-]][str2[j-]],
max(dp[i][j-] + score[][str2[j-]], dp[i-][j] + score[str1[i-]][]));
}
}
printf("%d\n", dp[len1][len2]);
} return ;
}
POJ1080(LCS变形)的更多相关文章
- poj 1080 (LCS变形)
Human Gene Functions 题意: LCS: 设dp[i][j]为前i,j的最长公共序列长度: dp[i][j] = dp[i-1][j-1]+1;(a[i] == b[j]) dp[i ...
- POJ 1080( LCS变形)
题目链接: http://poj.org/problem?id=1080 Human Gene Functions Time Limit: 1000MS Memory Limit: 10000K ...
- UVA-1625-Color Length(DP LCS变形)
Color Length(UVA-1625)(DP LCS变形) 题目大意 输入两个长度分别为n,m(<5000)的颜色序列.要求按顺序合成同一个序列,即每次可以把一个序列开头的颜色放到新序列的 ...
- Advanced Fruits(HDU 1503 LCS变形)
Advanced Fruits Time Limit: 2000/1000 MS (Java/Others) Memory Limit: 65536/32768 K (Java/Others)T ...
- poj1080--Human Gene Functions(dp:LCS变形)
Human Gene Functions Time Limit: 1000MS Memory Limit: 10000K Total Submissions: 17206 Accepted: ...
- HDU 5791 Two ——(LCS变形)
感觉就是最长公共子序列的一个变形(虽然我也没做过LCS啦= =). 转移方程见代码吧.这里有一个要说的地方,如果a[i] == a[j]的时候,为什么不需要像不等于的时候那样减去一个dp[i-1][j ...
- uva 10723 Cyborg Genes(LCS变形)
题目:http://acm.hust.edu.cn/vjudge/contest/view.action?cid=107450#problem/C 题意:输入两个字符串,找一个最短的串,使得输入的两个 ...
- Combine String---hdu5727 &&& Zipper(LCS变形)
题目链接:http://poj.org/problem?id=2192 http://acm.split.hdu.edu.cn/showproblem.php?pid=5707 http://acm. ...
- HUST 4681 String (DP LCS变形)
题目链接:http://acm.hdu.edu.cn/showproblem.php?pid=4681 题目大意:给定三个字符串A,B,C 求最长的串D,要求(1)D是A的字序列 (2)D是B的子序列 ...
随机推荐
- Gradle下载依赖jar包位置修改
gradle会下载相关需要依赖的jar包,默认的本地存放地址是:C:/Users/(用户名)/.gradle/caches/modules-2/files-2.1,很多人和我一样不愿意放在C盘,所以需 ...
- 玩玩自动化测试之selenium篇
现如今社会科技发展太快了,纯功能点点点已经落后别人好几条街了,所以为了让自己多点职业生涯年限,得挺起肩,傲起头.自动化测试,其本质是用代码程序测试程序,所以其实第一步应该学好编程语言,后再自己开发自动 ...
- json与python解析
1.json.dumps 将 Python 对象编码成 JSON 字符串 json.loads 将已编码的 JSON 字符串解码为 Python 对象 2.json.dump() ...
- deeplearning.ai课程学习(1)
本系列主要是我对吴恩达的deeplearning.ai课程的理解和记录,完整的课程笔记已经有很多了,因此只记录我认为重要的东西和自己的一些理解. 第一门课 神经网络和深度学习(Neural Netwo ...
- [译]在Python中如何使用额enumerate 和 zip 来迭代两个列表和它们的index?
enumerate - 迭代一个列表的index和item <Python Cookbook>(Recipe 4.4)描述了如何使用enumerate迭代item和index. 例子如下: ...
- 一个简单的NetCore项目:1 - 搭建框架,生成数据库
1- 启动项目 安装.NETCORE SDK,教程在网上可以搜索的到,这里就不讲述了.简单粗暴的方式就是安装最新的VS2015. 2-搭建框架 2.1 打开VS新建一个项目,在弹出的新建项目对话框中, ...
- Hash表 算法的详细解析
http://xingyunbaijunwei.blog.163.com/blog/static/76538067201111494524190/ 什么是HashHash,一般翻译做“散列”,也有直接 ...
- 《学习OpenCV》课后习题解答7
题目:(P105) 创建一个结构,结构中包含一个整数,一个CvPoint和一个 CvRect:称结构体为"my_struct". a. 写两个函数:void Write_my_st ...
- 201621044079WEEK06-接口、内部类
作业06-接口.内部类 1. 本周学习总结 1.1 面向对象学习暂告一段落,请使用思维导图,以封装.继承.多态为核心概念画一张思维导图或相关笔记,对面向对象思想进行一个总结. 注1:关键词与内容不求多 ...
- Java基础知识-去重
java基础知识-去掉list集合中的重复元素: 思路: 首先新建一个容器resultList用来存放去重之后的元素 然后遍历sourceList集合中的元素 判断所遍历的元素是否已经存在于resul ...
