Codeforces Round #295 (Div. 2)C - DNA Alignment 数学题
2 seconds
256 megabytes
standard input
standard output
Vasya became interested in bioinformatics. He's going to write an article about similar cyclic DNA sequences, so he invented a new method for determining the similarity of cyclic sequences.
Let's assume that strings s and t have the same length n, then the function h(s, t) is defined as the number of positions in which the respective symbols of s and t are the same. Function h(s, t) can be used to define the function of Vasya distance ρ(s, t):
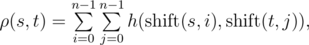 where
where 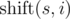 is obtained from string s, by applying left circular shift i times. For example,ρ("AGC", "CGT") = h("AGC", "CGT") + h("AGC", "GTC") + h("AGC", "TCG") + h("GCA", "CGT") + h("GCA", "GTC") + h("GCA", "TCG") + h("CAG", "CGT") + h("CAG", "GTC") + h("CAG", "TCG") = 1 + 1 + 0 + 0 + 1 + 1 + 1 + 0 + 1 = 6
is obtained from string s, by applying left circular shift i times. For example,ρ("AGC", "CGT") = h("AGC", "CGT") + h("AGC", "GTC") + h("AGC", "TCG") + h("GCA", "CGT") + h("GCA", "GTC") + h("GCA", "TCG") + h("CAG", "CGT") + h("CAG", "GTC") + h("CAG", "TCG") = 1 + 1 + 0 + 0 + 1 + 1 + 1 + 0 + 1 = 6
Vasya found a string s of length n on the Internet. Now he wants to count how many strings t there are such that the Vasya distance from the string s attains maximum possible value. Formally speaking, t must satisfy the equation: 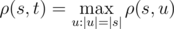 .
.
Vasya could not try all possible strings to find an answer, so he needs your help. As the answer may be very large, count the number of such strings modulo 109 + 7.
The first line of the input contains a single integer n (1 ≤ n ≤ 105).
The second line of the input contains a single string of length n, consisting of characters "ACGT".
Print a single number — the answer modulo 109 + 7.
1
C
1
2
AG
4
3
TTT
1
Please note that if for two distinct strings t1 and t2 values ρ(s, t1) и ρ(s, t2) are maximum among all possible t, then both strings must be taken into account in the answer even if one of them can be obtained by a circular shift of another one.
In the first sample, there is ρ("C", "C") = 1, for the remaining strings t of length 1 the value of ρ(s, t) is 0.
In the second sample, ρ("AG", "AG") = ρ("AG", "GA") = ρ("AG", "AA") = ρ("AG", "GG") = 4.
In the third sample, ρ("TTT", "TTT") = 27
int idx(char a)
{
if(a=='A')
return ;
if(a=='G')
return ;
if(a=='C')
return ;
if(a=='T')
return ;
}
int flag[];
int powmod(int a,int b) {
LL ans = ,x = a;
while (b) {
if (b & ) ans = ans * x % MOD;
x = x * x % MOD;
b >>= ;
}
return ans;
}
int main()
{
int n;
cin>>n;
string s;
cin>>s;
REP(i,s.size())
flag[idx(s[i])]++;
int max_num=;
int max_kiss=;
REP(i,)
max_num=max(max_num,flag[i]);
REP(i,)
{
if(flag[i]==max_num)
max_kiss++;
}
cout<<powmod(max_kiss,n)<<endl;
}
Codeforces Round #295 (Div. 2)C - DNA Alignment 数学题的更多相关文章
- codeforces 521a//DNA Alignment// Codeforces Round #295(Div. 1)
题意:如题定义的函数,取最大值的数量有多少? 结论只猜对了一半. 首先,如果只有一个元素结果肯定是1.否则.s串中元素数量分别记为a,t,c,g.设另一个串t中数量为a',t',c',g'.那么,固定 ...
- Codeforces Round #295 (Div. 2)
水 A. Pangram /* 水题 */ #include <cstdio> #include <iostream> #include <algorithm> # ...
- 【记忆化搜索】Codeforces Round #295 (Div. 2) B - Two Buttons
题意:给你一个数字n,有两种操作:减1或乘2,问最多经过几次操作能变成m: 随后发篇随笔普及下memset函数的初始化问题.自己也是涨了好多姿势. 代码 #include<iostream> ...
- Codeforces Round #295 (Div. 2)B - Two Buttons BFS
B. Two Buttons time limit per test 2 seconds memory limit per test 256 megabytes input standard inpu ...
- Codeforces Round #295 (Div. 2)A - Pangram 水题
A. Pangram time limit per test 2 seconds memory limit per test 256 megabytes input standard input ou ...
- Codeforces Round #295 (Div. 1) C. Pluses everywhere
昨天ZZD大神邀请我做一道题,说这题很有趣啊. 哇,然后我被虐了. Orz ZZD 题目大意: 你有一个长度为n的'0-9'串,你要在其中加入k个'+'号,每种方案就会形成一个算式,算式算出来的值记做 ...
- Codeforces Round #295 (Div. 2)---B. Two Buttons( bfs步数搜索记忆 )
B. Two Buttons time limit per test : 2 seconds memory limit per test :256 megabytes input :standard ...
- Codeforces Round #295 (Div. 2) B. Two Buttons
B. Two Buttons time limit per test 2 seconds memory limit per test 256 megabytes input standard inpu ...
- Codeforces Round #295 (Div. 2) B. Two Buttons 520B
B. Two Buttons time limit per test 2 seconds memory limit per test 256 megabytes input standard inpu ...
随机推荐
- mipi 调试经验【转】
转自:http://blog.csdn.net/g_salamander/article/details/9163455 版权声明:本文为博主原创文章,未经博主允许不得转载. 以下是最近几个月在调试 ...
- Tomcat安装与优化
Tomcat安装与优化 1.安装jdk环境 最新的JDK下载地址:http://www.oracle.com/technetwork/java/javase/downloads/jdk8-downlo ...
- mysql修改表的存储引擎(myisam<=>innodb)【转】
修改表的存储引擎myisam<=>innodb 查看表的存储引擎mysql> show create table tt7;+-------+--------------------- ...
- NuGet套件还原步骤(以vs2012为例)
下载别人的范例,出现由于Nuget套件不存在而无法启动时: 效果如下图: 步骤如下: 1.点击 项目->启用NuGet程序包还原 2.点击下图中的是 3.点击下图中的确定 4.效果如图: . 5 ...
- 九、springboot整合redis二之缓冲配置
1.创建Cache配置类 @Configuration @EnableCaching public class RedisCacheConfig extends CachingConfigurerSu ...
- 漂亮的SVG时钟
漂亮的SVG时钟 效果图: 代码如下,复制即可使用: <!DOCTYPE html> <html lang="en"> <head> <m ...
- ZooKeeper的典型应用场景
<从Paxos到Zookeeper 分布式一致性原理与实践>读书笔记 本文:总结脑图地址:脑图 前言 所有的典型应用场景,都是利用了ZK的如下特性: 强一致性:在高并发情况下,能够保证节点 ...
- SQLServer系统变量使用
1.@@IDENTITY返回最后插入的标识值.这个变量很有用,当你插入一行数据时,想同时获得该行的的ID(标示列),就可以用@@IDENTITY示例:下面的示例向带有标识列的表中插入一行,并用 @@I ...
- Spring介绍及配置(XML文件配置和注解配置)
本节内容: Spring介绍 Spring搭建 Spring概念 Spring配置讲解 使用注解配置Spring 一.Spring介绍 1. 什么是Spring Spring是一个开源框架,Sprin ...
- 一步一步学习IdentityServer4 (1) 概要配置说明
//结合EFCore生成IdentityServer4数据库 // 项目工程文件最后添加 <ItemGroup><DotNetCliToolReference Include=&qu ...
