POJ 1080 Human Gene Functions -- 动态规划(最长公共子序列)
题目地址:http://poj.org/problem?id=1080
Description
their functions, because these can be used to diagnose human diseases and to design new drugs for them.
A human gene can be identified through a series of time-consuming biological experiments, often with the help of computer programs. Once a sequence of a gene is obtained, the next job is to determine its function.
One of the methods for biologists to use in determining the function of a new gene sequence that they have just identified is to search a database with the new gene as a query. The database to be searched stores many gene sequences and their functions – many
researchers have been submitting their genes and functions to the database and the database is freely accessible through the Internet.
A database search will return a list of gene sequences from the database that are similar to the query gene.
Biologists assume that sequence similarity often implies functional similarity. So, the function of the new gene might be one of the functions that the genes from the list have. To exactly determine which one is the right one another series of biological experiments
will be needed.
Your job is to make a program that compares two genes and determines their similarity as explained below. Your program may be used as a part of the database search if you can provide an efficient one.
Given two genes AGTGATG and GTTAG, how similar are they? One of the methods to measure the similarity
of two genes is called alignment. In an alignment, spaces are inserted, if necessary, in appropriate positions of
the genes to make them equally long and score the resulting genes according to a scoring matrix.
For example, one space is inserted into AGTGATG to result in AGTGAT-G, and three spaces are inserted into GTTAG to result in –GT--TAG. A space is denoted by a minus sign (-). The two genes are now of equal
length. These two strings are aligned:
AGTGAT-G
-GT--TAG
In this alignment, there are four matches, namely, G in the second position, T in the third, T in the sixth, and G in the eighth. Each pair of aligned characters is assigned a score according to the following scoring matrix.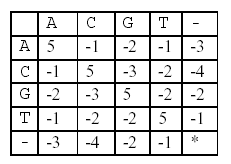
denotes that a space-space match is not allowed. The score of the alignment above is (-3)+5+5+(-2)+(-3)+5+(-3)+5=9.
Of course, many other alignments are possible. One is shown below (a different number of spaces are inserted into different positions):
AGTGATG
-GTTA-G
This alignment gives a score of (-3)+5+5+(-2)+5+(-1) +5=14. So, this one is better than the previous one. As a matter of fact, this one is optimal since no other alignment can have a higher score. So, it is said that the
similarity of the two genes is 14.
Input
The length of each gene sequence is at least one and does not exceed 100.
Output
Sample Input
2
7 AGTGATG
5 GTTAG
7 AGCTATT
9 AGCTTTAAA
Sample Output
14
21
#include <stdio.h>
int matrix[5][5] = {
{5, -1, -2, -1, -3},
{-1, 5, -3, -2, -4},
{-2, -3, 5, -2, -2},
{-1, -2, -2, 5, -1},
{-3, -4, -2, -1, 0},
};
char exchange[5] = {'A', 'C', 'G', 'T', ' '};
//通过矩阵求每一对字符的分值
int Value (char m, char n){
int R, C;
int i;
for (i=0; i<5; ++i){
if (exchange[i] == m)
R = i;
if (exchange[i] == n)
C = i;
}
return matrix[R][C];
}
int Max (int a, int b, int c){
int max = (a > b) ? a : b;
return (max > c) ? max : c;
}
int Similarity (char str1[], int length1, char str2[], int length2){
int dp[110][110];
int i, j;
dp[0][0] = 0;
for (i=1; i<=length1; ++i)
dp[i][0] = dp[i-1][0] + Value (str1[i], ' ');
for (i=1; i<=length2; ++i)
dp[0][i] = dp[0][i-1] + Value (' ', str2[i]);
///////////////////////////////////////////////////////////////////////////
//dp[i][j]表示第一个字串的长为i的子串与第二个字串的长为j的子串的相似度
for (i=1; i<=length1; ++i){
for (j=1; j<=length2; ++j){
///////////////////////////////////////////////////////////////////
//状态转移方程
if (str1[i] == str2[j]){
dp[i][j] = dp[i-1][j-1] + Value (str1[i], str2[j]);
}
else{
dp[i][j] = Max (dp[i-1][j] + Value (str1[i], ' '),
dp[i][j-1] + Value (' ', str2[j]),
dp[i-1][j-1] + Value (str1[i], str2[j]));
}
///////////////////////////////////////////////////////////////////
}
}
///////////////////////////////////////////////////////////////////////////
return dp[length1][length2];
}
int main(void){
char str1[110];
char str2[110];
int length1;
int length2;
int T;
int ans;
while (scanf ("%d", &T) != EOF){
while (T-- != 0){
scanf ("%d%s", &length1, str1 + 1);
scanf ("%d%s", &length2, str2 + 1);
if (length1 > length2)
ans = Similarity (str1, length1, str2, length2);
else
ans = Similarity (str2, length2, str1, length1);
printf ("%d\n", ans);
}
}
return 0;
}
HDOJ上相似的题目:http://acm.hdu.edu.cn/showproblem.php?pid=1513
POJ 1080 Human Gene Functions -- 动态规划(最长公共子序列)的更多相关文章
- poj 1080 ——Human Gene Functions——————【最长公共子序列变型题】
Human Gene Functions Time Limit: 1000MS Memory Limit: 10000K Total Submissions: 17805 Accepted: ...
- poj 1080 Human Gene Functions (最长公共子序列变形)
题意:有两个代表基因序列的字符串s1和s2,在两个基因序列中通过添加"-"来使得两个序列等长:其中每对基因匹配时会形成题中图片所示匹配值,求所能得到的总的最大匹配值. 题解:这题运 ...
- poj 1080 Human Gene Functions(lcs,较难)
Human Gene Functions Time Limit: 1000MS Memory Limit: 10000K Total Submissions: 19573 Accepted: ...
- dp poj 1080 Human Gene Functions
题目链接: http://poj.org/problem?id=1080 题目大意: 给两个由A.C.T.G四个字符组成的字符串,可以在两串中加入-,使得两串长度相等. 每两个字符匹配时都有个值,求怎 ...
- poj 1080 Human Gene Functions(dp)
题目:http://poj.org/problem?id=1080 题意:比较两个基因序列,测定它们的相似度,将两个基因排成直线,如果需要的话插入空格,使基因的长度相等,然后根据那个表格计算出相似度. ...
- POJ 1080 Human Gene Functions 【dp】
题目大意:每次给出两个碱基序列(包含ATGC的两个字符串),其中每一个碱基与另一串中碱基如果配对或者与空串对应会有一个分数(可能为负),找出一种方式使得两个序列配对的分数最大 思路:字符串动态规划的经 ...
- POJ 1080 Human Gene Functions
题意:给两个DNA序列,在这两个DNA序列中插入若干个'-',使两段序列长度相等,对应位置的两个符号的得分规则给出,求最高得分. 解法:dp.dp[i][j]表示第一个字符串s1的前i个字符和第二个字 ...
- HDU 1080 Human Gene Functions--DP--(变形最长公共子)
意甲冠军:该基因序列的两端相匹配,四种不同的核苷酸TCGA有不同的分值匹配.例如T-G比分是-2,它也可以被加入到空格,空洞格并且还具有一个相应的核苷酸匹配分值,求最大比分 分析: 在空气中的困难格的 ...
- POJ 1458 Common Subsequence(LCS最长公共子序列)
POJ 1458 Common Subsequence(LCS最长公共子序列)解题报告 题目链接:http://acm.hust.edu.cn/vjudge/contest/view.action?c ...
随机推荐
- jeewx的使用_01 接入和验证
jeewx是java语言的用于开发微信公共平台的一个框架,有人说很臃肿,但个人感觉还不错,仁者见仁智者见智吧, 下面简单介绍工作原理: 1.下载 要使用jeewx需要先下载其源码 jeewx介绍:ht ...
- 记一次js中和php中的字符串长度计算截取的终极问题和完美解决方案
1.js是用unicode算长度的,比如单字节的算1,中文也算1,但是正常我们想让两个单字节算1,如何计算这个长度 第一种解决方案,用正则,如下 /[\u0x00-\u0xff]/,天真的想着,这样就 ...
- 辛星浅谈PHP的混乱的编码风格
我们都知道.各种编程语言都有自己的风格,即使是像C和C++那样一脉相承的语言(C++本意全然兼容C的语法).编程风格上还是有些区别.比方非常典型的就是C++风格的单行凝视和C风格的多行凝视. 而尽管J ...
- 【M12】了解“抛出一个exception”与“传递一个参数”或“调用一个虚函数”之间的差异
1.方法参数的声明语法和catch语句的语法是一样的,你可能会认为主调方法调用一个方法,并向其传递参数,与抛出一个异常传递到catch语句是一样的,是的,有相同之处,但也有更大的不同. 2.主调方法调 ...
- 部署SharePoint解决方式包时遇到的问题
部署SharePoint解决方式包时遇到的问题 近期我在使用STSADM.EXE命令部署解决方式包的时候.遇到一个问题.很的难搞. 创建WSP文件非常easy.加入到解决方式库也非常e ...
- Android Intent入门
http://www.cnblogs.com/leipei2352/archive/2011/08/09/2132096.html http://blog.csdn.net/xiazdong/arti ...
- Android手机监控软件设计实现
一.需求分析: 随着IT信息技术的飞速发展,手机的普及,伴随着智能手机的出现及快速的更新换代,手机已不仅仅是一个通信工具,更是一个多功能的应用平台. 手机监控软件则是基于电脑监控软件的原理,植入手机平 ...
- SQL Server日期时间格式转换字符串详解
本文我们主要介绍了SQL Server日期时间格式转换字符串的相关知识,并给出了大量实例对其各个参数进行对比说明,希望能够对您有所帮助. 在SQL Server数据库中,SQL Server日期时间格 ...
- 同时使用ADO与Excel类库冲突的问题
客户需要一个Demo程序实现Access数据库表导出到Excel表格,并将表中存储的照片(OLE对象)以其中一个字段(编号)命名存储到本地.程序中引入了ADO操作Access数据库("C:\ ...
- 101个直接可以拿来用的JavaScript实用功能代码片段(转)
1.原生JavaScript实现字符串长度截取 function cutstr(str, len) { var temp; var icount = 0; var patrn = /[^x00-xff ...
