Best packages for data manipulation in R
dplyr and data.table are amazing packages that make data manipulation in R fun. Both packages have their strengths. While dplyr is more elegant and resembles natural language, data.table is succinct and we can do a lot withdata.table in just a single line. Further, data.table is, in some cases, faster (see benchmark here) and it may be a go-to package when performance and memory are constraints. You can read comparison of dplyr and data.tablefrom Stack Overflow and Quora.
You can get reference manual and vignettes for data.table here and for dplyrhere. You can read other tutorial about dplyr published at DataScience+
Background
I am a long time dplyr and data.table user for my data manipulation tasks. For someone who knows one of these packages, I thought it could help to show codes that perform the same tasks in both packages to help them quickly study the other. If you know either package and have interest to study the other, this post is for you.
dplyr
dplyr has 5 verbs which make up the majority of the data manipulation tasks we perform. Select: used to select one or more columns; Filter: used to select some rows based on specific criteria; Arrange: used to sort data based on one or more columns in ascending or descending order; Mutate: used to add new columns to our data; Summarise: used to create chunks from our data.
data.table
data.table has a very succinct general format: DT[i, j, by], which is interpreted as: Take DT, subset rows using i, then calculate j grouped by by.
Data manipulation
First we will install some packages for our project.
library(dplyr)
library(data.table)
library(lubridate)
library(jsonlite)
library(tidyr)
library(ggplot2)
library(compare)
The data we will use here is from DATA.GOV. It is Medicare Hospital Spending by Claim and it can be downloaded from here. Let’s download the data in JSONformat using the fromJSON function from the jsonlite package. Since JSON is a very common data format used for asynchronous browser/server communication, it is good if you understand the lines of code below used to get the data. You can get an introductory tutorial on how to use the jsonlite package to work with JSON data here and here. However, if you want to focus only on the data.table and dplyr commands, you can safely just run the codes in the two cells below and ignore the details.
spending=fromJSON("https://data.medicare.gov/api/views/nrth-mfg3/rows.json?accessType=DOWNLOAD")
names(spending)
"meta" "data"
meta=spending$meta
hospital_spending=data.frame(spending$data)
colnames(hospital_spending)=make.names(meta$view$columns$name)
hospital_spending=select(hospital_spending,-c(sid:meta))
glimpse(hospital_spending)
Observations: 70598
Variables:
$ Hospital.Name (fctr) SOUTHEAST ALABAMA MEDICAL CENT...
$ Provider.Number. (fctr) 010001, 010001, 010001, 010001...
$ State (fctr) AL, AL, AL, AL, AL, AL, AL, AL...
$ Period (fctr) 1 to 3 days Prior to Index Hos...
$ Claim.Type (fctr) Home Health Agency, Hospice, I...
$ Avg.Spending.Per.Episode..Hospital. (fctr) 12, 1, 6, 160, 1, 6, 462, 0, 0...
$ Avg.Spending.Per.Episode..State. (fctr) 14, 1, 6, 85, 2, 9, 492, 0, 0,...
$ Avg.Spending.Per.Episode..Nation. (fctr) 13, 1, 5, 117, 2, 9, 532, 0, 0...
$ Percent.of.Spending..Hospital. (fctr) 0.06, 0.01, 0.03, 0.84, 0.01, ...
$ Percent.of.Spending..State. (fctr) 0.07, 0.01, 0.03, 0.46, 0.01, ...
$ Percent.of.Spending..Nation. (fctr) 0.07, 0.00, 0.03, 0.58, 0.01, ...
$ Measure.Start.Date (fctr) 2014-01-01T00:00:00, 2014-01-0...
$ Measure.End.Date (fctr) 2014-12-31T00:00:00, 2014-12-3...
As shown above, all columns are imported as factors and let’s change the columns that contain numeric values to numeric.
cols = 6:11; # These are the columns to be changed to numeric.
hospital_spending[,cols] <- lapply(hospital_spending[,cols], as.numeric)
The last two columns are measure start date and measure end date. So, let’s use the lubridate package to correct the classes of these columns.
cols = 12:13; # These are the columns to be changed to dates.
hospital_spending[,cols] <- lapply(hospital_spending[,cols], ymd_hms)
Now, let’s check if the columns have the classes we want.
sapply(hospital_spending, class)
$Hospital.Name
"factor"
$Provider.Number.
"factor"
$State
"factor"
$Period
"factor"
$Claim.Type
"factor"
$Avg.Spending.Per.Episode..Hospital.
"numeric"
$Avg.Spending.Per.Episode..State.
"numeric"
$Avg.Spending.Per.Episode..Nation.
"numeric"
$Percent.of.Spending..Hospital.
"numeric"
$Percent.of.Spending..State.
"numeric"
$Percent.of.Spending..Nation.
"numeric"
$Measure.Start.Date
"POSIXct" "POSIXt"
$Measure.End.Date
"POSIXct" "POSIXt"
Create data table
We can create a data.table using the data.table() function.
hospital_spending_DT = data.table(hospital_spending)
class(hospital_spending_DT)
"data.table" "data.frame"
Select certain columns of data
To select columns, we use the verb select in dplyr. In data.table, on the other hand, we can specify the column names.
Selecting one variable
Let’s selet the “Hospital Name” variable
from_dplyr = select(hospital_spending, Hospital.Name)
from_data_table = hospital_spending_DT[,.(Hospital.Name)]
Now, let’s compare if the results from dplyr and data.table are the same.
compare(from_dplyr,from_data_table, allowAll=TRUE)
TRUE
dropped attributes
Removing one variable
from_dplyr = select(hospital_spending, -Hospital.Name)
from_data_table = hospital_spending_DT[,!c("Hospital.Name"),with=FALSE]
compare(from_dplyr,from_data_table, allowAll=TRUE)
TRUE
dropped attributes
we can also use := function which modifies the input data.table by reference.
We will use the copy() function, which deep copies the input object and therefore any subsequent update by reference operations performed on the copied object will not affect the original object.
DT=copy(hospital_spending_DT)
DT=DT[,Hospital.Name:=NULL]
"Hospital.Name"%in%names(DT)FALSE
We can also remove many variables at once similarly:
DT=copy(hospital_spending_DT)
DT=DT[,c("Hospital.Name","State","Measure.Start.Date","Measure.End.Date"):=NULL]
c("Hospital.Name","State","Measure.Start.Date","Measure.End.Date")%in%names(DT)
FALSE FALSE FALSE FALSE
Selecting multiple variables
Let’s select the variables:
Hospital.Name,State,Measure.Start.Date,and Measure.End.Date.
from_dplyr = select(hospital_spending, Hospital.Name,State,Measure.Start.Date,Measure.End.Date)
from_data_table = hospital_spending_DT[,.(Hospital.Name,State,Measure.Start.Date,Measure.End.Date)]
compare(from_dplyr,from_data_table, allowAll=TRUE)
TRUE
dropped attributes
Dropping multiple variables
Now, let’s remove the variables Hospital.Name,State,Measure.Start.Date,and Measure.End.Date from the original data frame hospital_spending and the data.table hospital_spending_DT.
from_dplyr = select(hospital_spending, -c(Hospital.Name,State,Measure.Start.Date,Measure.End.Date))
from_data_table = hospital_spending_DT[,!c("Hospital.Name","State","Measure.Start.Date","Measure.End.Date"),with=FALSE]
compare(from_dplyr,from_data_table, allowAll=TRUE)
TRUE
dropped attributes
dplyr has functions contains(), starts_with() and, ends_with() which we can use with the verb select. In data.table, we can use regular expressions. Let’s select columns that contain the word Date to demonstrate by example.
from_dplyr = select(hospital_spending,contains("Date"))
from_data_table = subset(hospital_spending_DT,select=grep("Date",names(hospital_spending_DT)))
compare(from_dplyr,from_data_table, allowAll=TRUE)
TRUE
dropped attributes
names(from_dplyr)
"Measure.Start.Date" "Measure.End.Date"
Rename columns
setnames(hospital_spending_DT,c("Hospital.Name", "Measure.Start.Date","Measure.End.Date"), c("Hospital","Start_Date","End_Date"))
names(hospital_spending_DT)
"Hospital" "Provider.Number." "State" "Period" "Claim.Type" "Avg.Spending.Per.Episode..Hospital." "Avg.Spending.Per.Episode..State." "Avg.Spending.Per.Episode..Nation." "Percent.of.Spending..Hospital." "Percent.of.Spending..State." "Percent.of.Spending..Nation." "Start_Date" "End_Date"
hospital_spending = rename(hospital_spending,Hospital= Hospital.Name, Start_Date=Measure.Start.Date,End_Date=Measure.End.Date)
compare(hospital_spending,hospital_spending_DT, allowAll=TRUE)
TRUE
dropped attributes
Filtering data to select certain rows
To filter data to select specific rows, we use the verb filter from dplyr with logical statements that could include regular expressions. In data.table, we need the logical statements only.
Filter based on one variable
from_dplyr = filter(hospital_spending,State=='CA') # selecting rows for California
from_data_table = hospital_spending_DT[State=='CA']
compare(from_dplyr,from_data_table, allowAll=TRUE)
TRUE
dropped attributes
Filter based on multiple variables
from_dplyr = filter(hospital_spending,State=='CA' & Claim.Type!="Hospice")
from_data_table = hospital_spending_DT[State=='CA' & Claim.Type!="Hospice"]
compare(from_dplyr,from_data_table, allowAll=TRUE)
TRUE
dropped attributes
from_dplyr = filter(hospital_spending,State %in% c('CA','MA',"TX"))
from_data_table = hospital_spending_DT[State %in% c('CA','MA',"TX")]
unique(from_dplyr$State)
CA MA TX
compare(from_dplyr,from_data_table, allowAll=TRUE)
TRUE
dropped attributes
Order data
We use the verb arrange in dplyr to order the rows of data. We can order the rows by one or more variables. If we want descending, we have to use desc()as shown in the examples.The examples are self-explanatory on how to sort in ascending and descending order. Let’s sort using one variable.
Ascending
from_dplyr = arrange(hospital_spending, State)
from_data_table = setorder(hospital_spending_DT, State)
compare(from_dplyr,from_data_table, allowAll=TRUE)
TRUE
dropped attributes
Descending
from_dplyr = arrange(hospital_spending, desc(State))
from_data_table = setorder(hospital_spending_DT, -State)
compare(from_dplyr,from_data_table, allowAll=TRUE)
TRUE
dropped attributes
Sorting with multiple variables
Let’s sort with State in ascending order and End_Date in descending order.
from_dplyr = arrange(hospital_spending, State,desc(End_Date))
from_data_table = setorder(hospital_spending_DT, State,-End_Date)
compare(from_dplyr,from_data_table, allowAll=TRUE)
TRUE
dropped attributes
Adding/updating column(s)
In dplyr we use the function mutate() to add columns. In data.table, we can Add/update a column by reference using := in one line.
from_dplyr = mutate(hospital_spending, diff=Avg.Spending.Per.Episode..State. - Avg.Spending.Per.Episode..Nation.)
from_data_table = copy(hospital_spending_DT)
from_data_table = from_data_table[,diff := Avg.Spending.Per.Episode..State. - Avg.Spending.Per.Episode..Nation.]
compare(from_dplyr,from_data_table, allowAll=TRUE)
TRUE
sorted
renamed rows
dropped row names
dropped attributes
from_dplyr = mutate(hospital_spending, diff1=Avg.Spending.Per.Episode..State. - Avg.Spending.Per.Episode..Nation.,diff2=End_Date-Start_Date)
from_data_table = copy(hospital_spending_DT)
from_data_table = from_data_table[,c("diff1","diff2") := list(Avg.Spending.Per.Episode..State. - Avg.Spending.Per.Episode..Nation.,diff2=End_Date-Start_Date)]
compare(from_dplyr,from_data_table, allowAll=TRUE)
TRUE
dropped attributes
Summarizing columns
We can use the summarize() function from dplyr to create summary statistics.
summarize(hospital_spending,mean=mean(Avg.Spending.Per.Episode..Nation.))
mean 8.772727 hospital_spending_DT[,.(mean=mean(Avg.Spending.Per.Episode..Nation.))]
mean 8.772727 summarize(hospital_spending,mean=mean(Avg.Spending.Per.Episode..Nation.),
maximum=max(Avg.Spending.Per.Episode..Nation.),
minimum=min(Avg.Spending.Per.Episode..Nation.),
median=median(Avg.Spending.Per.Episode..Nation.))
mean maximum minimum median
8.77 19 1 8.5 hospital_spending_DT[,.(mean=mean(Avg.Spending.Per.Episode..Nation.),
maximum=max(Avg.Spending.Per.Episode..Nation.),
minimum=min(Avg.Spending.Per.Episode..Nation.),
median=median(Avg.Spending.Per.Episode..Nation.))]
mean maximum minimum median
8.77 19 1 8.5
We can calculate our summary statistics for some chunks separately. We use the function group_by() in dplyr and in data.table, we simply provide by.
head(hospital_spending_DT[,.(mean=mean(Avg.Spending.Per.Episode..Hospital.)),by=.(Hospital)])
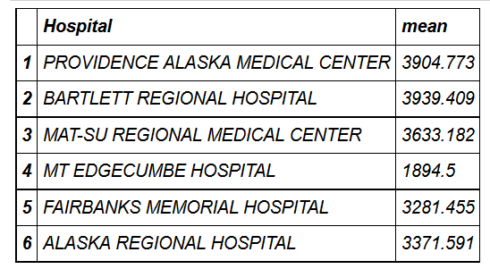
mygroup= group_by(hospital_spending,Hospital)
from_dplyr = summarize(mygroup,mean=mean(Avg.Spending.Per.Episode..Hospital.))
from_data_table=hospital_spending_DT[,.(mean=mean(Avg.Spending.Per.Episode..Hospital.)), by=.(Hospital)]
compare(from_dplyr,from_data_table, allowAll=TRUE) TRUE
sorted
renamed rows
dropped row names
dropped attributes
We can also provide more than one grouping condition.
head(hospital_spending_DT[,.(mean=mean(Avg.Spending.Per.Episode..Hospital.)),
by=.(Hospital,State)])
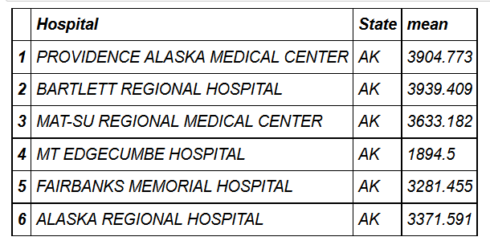
mygroup= group_by(hospital_spending,Hospital,State)
from_dplyr = summarize(mygroup,mean=mean(Avg.Spending.Per.Episode..Hospital.))
from_data_table=hospital_spending_DT[,.(mean=mean(Avg.Spending.Per.Episode..Hospital.)), by=.(Hospital,State)]
compare(from_dplyr,from_data_table, allowAll=TRUE)
TRUE
sorted
renamed rows
dropped row names
dropped attributes
Chaining
With both dplyr and data.table, we can chain functions in succession. In dplyr, we use pipes from the magrittr package with %>% which is really cool. %>% takes the output from one function and feeds it to the first argument of the next function. In data.table, we can use %>% or [ for chaining.
from_dplyr=hospital_spending%>%group_by(Hospital,State)%>%summarize(mean=mean(Avg.Spending.Per.Episode..Hospital.))
from_data_table=hospital_spending_DT[,.(mean=mean(Avg.Spending.Per.Episode..Hospital.)), by=.(Hospital,State)]
compare(from_dplyr,from_data_table, allowAll=TRUE)
TRUE
sorted
renamed rows
dropped row names
dropped attributes
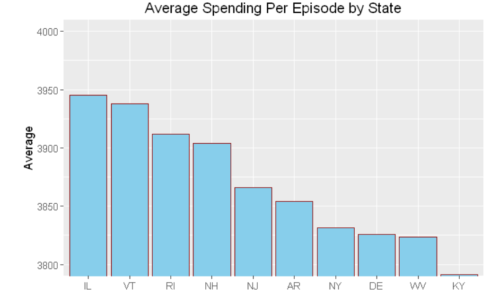
hospital_spending%>%group_by(State)%>%summarize(mean=mean(Avg.Spending.Per.Episode..Hospital.))%>%
arrange(desc(mean))%>%head(10)%>%
mutate(State = factor(State,levels = State[order(mean,decreasing =TRUE)]))%>%
ggplot(aes(x=State,y=mean))+geom_bar(stat='identity',color='darkred',fill='skyblue')+
xlab("")+ggtitle('Average Spending Per Episode by State')+
ylab('Average')+ coord_cartesian(ylim = c(3800, 4000))
hospital_spending_DT[,.(mean=mean(Avg.Spending.Per.Episode..Hospital.)),
by=.(State)][order(-mean)][1:10]%>%
mutate(State = factor(State,levels = State[order(mean,decreasing =TRUE)]))%>%
ggplot(aes(x=State,y=mean))+geom_bar(stat='identity',color='darkred',fill='skyblue')+
xlab("")+ggtitle('Average Spending Per Episode by State')+
ylab('Average')+ coord_cartesian(ylim = c(3800, 4000))

Summary
In this blog post, we saw how we can perform the same tasks using data.tableand dplyr packages. Both packages have their strengths. While dplyr is more elegant and resembles natural language, data.table is succinct and we can do a lot with data.table in just a single line. Further, data.table is, in some cases, faster and it may be a go-to package when performance and memory are the constraints.
You can get the code for this blog post at my GitHub account.
This is enough for this post. If you have any questions or feedback, feel free to leave a comment.
转自:http://datascienceplus.com/best-packages-for-data-manipulation-in-r/
Best packages for data manipulation in R的更多相关文章
- Data manipulation primitives in R and Python
Data manipulation primitives in R and Python Both R and Python are incredibly good tools to manipula ...
- Data Manipulation with dplyr in R
目录 select The filter and arrange verbs arrange filter Filtering and arranging Mutate The count verb ...
- The dplyr package has been updated with new data manipulation commands for filters, joins and set operations.(转)
dplyr 0.4.0 January 9, 2015 in Uncategorized I’m very pleased to announce that dplyr 0.4.0 is now av ...
- An Introduction to Stock Market Data Analysis with R (Part 1)
Around September of 2016 I wrote two articles on using Python for accessing, visualizing, and evalua ...
- 7 Tools for Data Visualization in R, Python, and Julia
7 Tools for Data Visualization in R, Python, and Julia Last week, some examples of creating visualiz ...
- java.sql.SQLException: Can not issue data manipulation statements with executeQuery().
1.错误描写叙述 java.sql.SQLException: Can not issue data manipulation statements with executeQuery(). at c ...
- Can not issue data manipulation statements with executeQuery()错误解决
转: Can not issue data manipulation statements with executeQuery()错误解决 2012年03月27日 15:47:52 katalya 阅 ...
- 数据库原理及应用-SQL数据操纵语言(Data Manipulation Language)和嵌入式SQL&存储过程
2018-02-19 18:03:54 一.数据操纵语言(Data Manipulation Language) 数据操纵语言是指插入,删除和更新语言. 二.视图(View) 数据库三级模式,两级映射 ...
- Can not issue data manipulation statements with executeQuery().解决方案
这个错误提示是说无法发行sql语句到指定的位置 错误写法: 正确写法: excuteQuery是查询语句,而我要调用的是更新的语句,所以这样数据库很为难到底要干嘛,实际我想用的是更新,但是我写成了查询 ...
随机推荐
- package(1):tm
tm包是R语言中为文本挖掘提供综合性处理的package,进行操作前载入tm包,vignette命令可以让你得到相关的文档说明.使用默认安装的R平台是不带tm package的,在安装的过程中,它会 ...
- QQ_MultiTalkServer
package test_teacher;import java.net.*;import java.io.*;public class MultiTalkServer{ public stat ...
- 在ASP.NET Core中使用Apworks开发数据服务:对HAL的支持
HAL,全称为Hypertext Application Language,它是一种简单的数据格式,它能以一种简单.统一的形式,在API中引入超链接特性,使得API的可发现性(discoverable ...
- MySql字符串函数使用技巧
1.从左开始截取字符串 left(str, length) 说明:left(被截取字段,截取长度) 例:select left(content,200) as abstract from my_con ...
- 初学Java scirpt(判断、循环语句)
在编写代码时,我们经常需要为不同的判断结果来执行不同的动作以及需要反复执行同一段代码,这时我们就需要使用判断和循环语句来实现. 1.判断语句(if) 判断语句经常用的有(if......else).( ...
- [原创] JavaScript实现简单的颜色类标签云
效果预览: 源码分享: <!DOCTYPE html><html><head lang="en"> <meta charset=" ...
- LVS + keepalived(DR) 实战
一.LVS体系结构 使用LVS架设的服务器集群系统有三个部分组成:最前端的负载均衡层,用Load Balancer表示,中间的服务器群组层,用Server Array表示,最底端的数据共享存储层,用S ...
- html5表单元素详解
表单是Html中获取用户输入的手段.此文对表单的元素进行了详细整理. 表单基本元素 form input button form元素 html4中,form元素相当于表单的外包装,其他都要在里面.ht ...
- 在线恶意软件和URL分析集成框架 – MalSub
malsub是一个基于Python 3.6.x的框架,它的设计遵循了当前最流行的互联网软件架构RESTful架构,并通过其RESTful API应用程序编程接口(API),封装了多个在线恶意软件和UR ...
- 清北Day 2
清北第二天,感受到了来自这个世界的不友善,大概把没听过不会的"名词"记录下来就已经一面了,然后被大佬说这都是最基础的东西,就很皮,那就趁别人练习字符串的题的时候,来写波博客了,倒不 ...
