More 3D Graphics (rgl) for Classification with Local Logistic Regression and Kernel Density Estimates (from The Elements of Statistical Learning)(转)
This post builds on a previous post, but can be read and understood independently.
As part of my course on statistical learning, we created 3D graphics to foster a more intuitive understanding of the various methods that are used to relax the assumption of linearity (in the predictors) in regression and classification methods.
The authors of our text (The Elements of Statistical Learning, 2nd Edition) provide a Mixture Simulation data set that has two continuous predictors and a binary outcome. This data is used to demonstrate classification procedures by plotting classification boundaries in the two predictors, which are determined by one or more surfaces (e.g., a probability surface such as that produced by logistic regression, or multiple intersecting surfaces as in linear discriminant analysis). In our class laboratory, we used the R package rgl to create a 3D representation of these surfaces for a variety of semiparametric classification procedures.
Chapter 6 presents local logistic regression and kernel density classification, among other kernel (local) classification and regression methods. Below is the code and graphic (a 2D projection) associated with the local linear logistic regression in these data:
library(rgl)
load(url("http://statweb.stanford.edu/~tibs/ElemStatLearn/datasets/ESL.mixture.rda"))
dat <- ESL.mixture
ddat <- data.frame(y=dat$y, x1=dat$x[,1], x2=dat$x[,2]) ## create 3D graphic, rotate to view 2D x1/x2 projection
par3d(FOV=1,userMatrix=diag(4))
plot3d(dat$xnew[,1], dat$xnew[,2], dat$prob, type="n",
xlab="x1", ylab="x2", zlab="",
axes=FALSE, box=TRUE, aspect=1) ## plot points and bounding box
x1r <- range(dat$px1)
x2r <- range(dat$px2)
pts <- plot3d(dat$x[,1], dat$x[,2], 1,
type="p", radius=0.5, add=TRUE,
col=ifelse(dat$y, "orange", "blue"))
lns <- lines3d(x1r[c(1,2,2,1,1)], x2r[c(1,1,2,2,1)], 1) ## draw Bayes (True) classification boundary in blue
dat$probm <- with(dat, matrix(prob, length(px1), length(px2)))
dat$cls <- with(dat, contourLines(px1, px2, probm, levels=0.5))
pls0 <- lapply(dat$cls, function(p) lines3d(p$x, p$y, z=1, color="blue")) ## compute probabilities plot classification boundary
## associated with local linear logistic regression
probs.loc <-
apply(dat$xnew, 1, function(x0) {
## smoothing parameter
l <- 1/2
## compute (Gaussian) kernel weights
d <- colSums((rbind(ddat$x1, ddat$x2) - x0)^2)
k <- exp(-d/2/l^2)
## local fit at x0
fit <- suppressWarnings(glm(y ~ x1 + x2, data=ddat, weights=k,
family=binomial(link="logit")))
## predict at x0
as.numeric(predict(fit, type="response", newdata=as.data.frame(t(x0))))
}) dat$probm.loc <- with(dat, matrix(probs.loc, length(px1), length(px2)))
dat$cls.loc <- with(dat, contourLines(px1, px2, probm.loc, levels=0.5))
pls <- lapply(dat$cls.loc, function(p) lines3d(p$x, p$y, z=1)) ## plot probability surface and decision plane
sfc <- surface3d(dat$px1, dat$px2, probs.loc, alpha=1.0,
color="gray", specular="gray")
qds <- quads3d(x1r[c(1,2,2,1)], x2r[c(1,1,2,2)], 0.5, alpha=0.4,
color="gray", lit=FALSE)
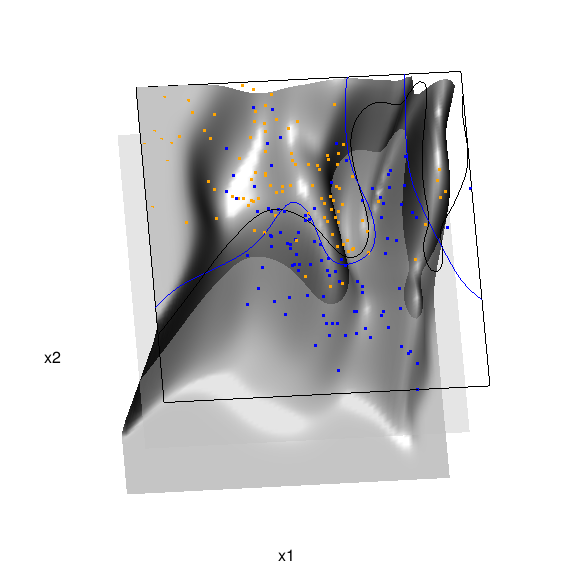
In the above graphic, the solid blue line represents the true Bayes decision boundary (i.e., {x: Pr("orange"|x) = 0.5}), which is computed from the model used to simulate these data. The probability surface (generated by the local logistic regression) is represented in gray, and the corresponding Bayes decision boundary occurs where the plane f(x) = 0.5 (in light gray) intersects with the probability surface. The solid black line is a projection of this intersection. Here is a link to the interactive version of this graphic: local logistic regression.
Below is the code and graphic associated with the kernel density classification (note: this code below should only be executed after the above code, since the 3D graphic is modified, rather than created anew):
## clear the surface, decision plane, and decision boundary
pop3d(id=sfc); pop3d(id=qds)
for(pl in pls) pop3d(id=pl) ## kernel density classification
## compute kernel density estimates for each class
dens.kde <-
lapply(unique(ddat$y), function(uy) {
apply(dat$xnew, 1, function(x0) {
## subset to current class
dsub <- subset(ddat, y==uy)
## smoothing parameter
l <- 1/2
## kernel density estimate at x0
mean(dnorm(dsub$x1-x0[1], 0, l)*dnorm(dsub$x2-x0[2], 0, l))
})
}) ## compute prior for each class (sample proportion)
prir.kde <- table(ddat$y)/length(dat$y) ## compute posterior probability Pr(y=1|x)
probs.kde <- prir.kde[2]*dens.kde[[2]]/
(prir.kde[1]*dens.kde[[1]]+prir.kde[2]*dens.kde[[2]]) ## plot classification boundary associated
## with kernel density classification
dat$probm.kde <- with(dat, matrix(probs.kde, length(px1), length(px2)))
dat$cls.kde <- with(dat, contourLines(px1, px2, probm.kde, levels=0.5))
pls <- lapply(dat$cls.kde, function(p) lines3d(p$x, p$y, z=1)) ## plot probability surface and decision plane
sfc <- surface3d(dat$px1, dat$px2, probs.kde, alpha=1.0,
color="gray", specular="gray")
qds <- quads3d(x1r[c(1,2,2,1)], x2r[c(1,1,2,2)], 0.5, alpha=0.4,
color="gray", lit=FALSE)
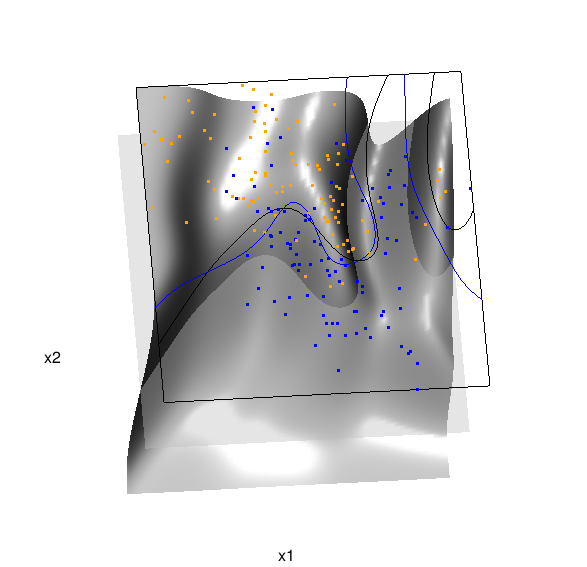
Here are links to the interactive versions of both graphics: local logistic regression, kernel density classification
This entry was posted in Technical and tagged data, graphics, programming, R, statistics on February 7, 2015.
More 3D Graphics (rgl) for Classification with Local Logistic Regression and Kernel Density Estimates (from The Elements of Statistical Learning)(转)的更多相关文章
- Some 3D Graphics (rgl) for Classification with Splines and Logistic Regression (from The Elements of Statistical Learning)(转)
This semester I'm teaching from Hastie, Tibshirani, and Friedman's book, The Elements of Statistical ...
- 李宏毅机器学习笔记3:Classification、Logistic Regression
李宏毅老师的机器学习课程和吴恩达老师的机器学习课程都是都是ML和DL非常好的入门资料,在YouTube.网易云课堂.B站都能观看到相应的课程视频,接下来这一系列的博客我都将记录老师上课的笔记以及自己对 ...
- Logistic Regression Using Gradient Descent -- Binary Classification 代码实现
1. 原理 Cost function Theta 2. Python # -*- coding:utf8 -*- import numpy as np import matplotlib.pyplo ...
- Classification week2: logistic regression classifier 笔记
华盛顿大学 machine learning: Classification 笔记. linear classifier 线性分类器 多项式: Logistic regression & 概率 ...
- Classification and logistic regression
logistic 回归 1.问题: 在上面讨论回归问题时.讨论的结果都是连续类型.但假设要求做分类呢?即讨论结果为离散型的值. 2.解答: 假设: 当中: g(z)的图形例如以下: 由此可知:当hθ( ...
- Android Programming 3D Graphics with OpenGL ES (Including Nehe's Port)
https://www3.ntu.edu.sg/home/ehchua/programming/android/Android_3D.html
- Logistic Regression and Classification
分类(Classification)与回归都属于监督学习,两者的唯一区别在于,前者要预测的输出变量\(y\)只能取离散值,而后者的输出变量是连续的.这些离散的输出变量在分类问题中通常称之为标签(Lab ...
- Logistic Regression求解classification问题
classification问题和regression问题类似,区别在于y值是一个离散值,例如binary classification,y值只取0或1. 方法来自Andrew Ng的Machine ...
- 分类和逻辑回归(Classification and logistic regression)
分类问题和线性回归问题问题很像,只是在分类问题中,我们预测的y值包含在一个小的离散数据集里.首先,认识一下二元分类(binary classification),在二元分类中,y的取值只能是0和1.例 ...
随机推荐
- 刷机无法连接4g
只显示2g,gsm only 无法修改,本人刷cm13和lineageOs都遇到过这样的情况,可能与手机有关xt1570(moto x style),特在此分享,希望有用 1.首先在设置中将sim卡网 ...
- HTML5技术实现Web图形图像处理——WebPhotoshop精简版
WebPhotoshop精简版是利用HTML5技术在Web上实现对图形图像的处理,构建易维护.易共享.易于拓展.实时性的Web图形图像处理平台. 精简版功能包括:图形绘制.图像处理.图像操作.完整版包 ...
- Android -- 从源码带你从EventBus2.0飚到EventBus3.0(一)
1,最近看了不少的面试题,不管是百度.网易.阿里的面试题,都会问到EventBus源码和RxJava源码,而自己只是在项目中使用过,却没有去用心的了解它底层是怎么实现的,所以今天就和大家一起来学习学习 ...
- dubbo个人总结
dubbo,分布式服务框架,RPC服务框架. 注册中心zk,redis......,使用大多为zk 注册流程 最后一图 服务提供者启动时向/dubbo/com.foo.BarService/provi ...
- 分享一本书<<谁都不敢欺负你>>
有些人,不管在工作还是生活上,总是被人欺负. 分享这本书给大家,能给大家带来正能量.你强大了,就没人敢欺负你. 有的时候,感到为什么倒霉的总是我?为什么我的命运是这样?为什么总欺负我? 也许有很多人会 ...
- Elasticsearch搜索之best_fields分析
顾名思义,best_field就是获取最佳匹配的field,另个可以通过tie_breaker来控制其他field的得分,boost可以设置权重(默认都为1). 下面从宏观上来讲的简单公式: scor ...
- uc广告过滤你能更坑点不
背景: 搞的手机站要上线,电脑测试木有问题,拿手机访问,有个页面始终不正常, 其他的 windows phone 的正常, ios 的也正常 就唯独 ,用的是安卓,uc的浏览器显示有问题 我勒个去,那 ...
- 跟着刚哥梳理java知识点——反射和代理(十七)
反射机制是什么?反射机制是在运行状态中,对于任意一个类,都能够知道这个类的所有的属性和方法:对于任意一个对象,都能够调用他的一个方法和属性,这种动态获取的信息以及动态调用对象的方法的功能称为java语 ...
- 【2017-04-24】winform基础、登录窗口、窗口属性
一.winform基础 客户端应用程序:C/S 客户端应用程序可以操作用户电脑中的文件,代码要在用户电脑上执行,吃用户电脑配置. 窗体是由控件和属性做出来的 控件:窗体里所放的东西."视图 ...
- 1.6 OWIN集成
OWIN集成 安装 使用 如果在应用程序里既使用ASP.NET MVC也使用ASP.NET Web API,需要在工程里安装Abp.Owin包. 安装 添加Abp.Owin包到主工程里(一般是web工 ...
