Tree - Gradient Boosting Machine with sklearn source code
This is the second post in Boosting algorithm. In the previous post, we go through the earliest Boosting algorithm - AdaBoost, which is actually an approximation of exponential loss via additive stage-forward modelling. What if we want to choose other loss function? Can we have a more generic algorithm that can apply to all loss function.
Gradient Descent
Friedman proposed another way to optimize the additive function- Gradient descent, same as the numerical optimization method used in neural network. Basically at each step we calculate the gradient against current additive function \(F_m(x)\) to first find the direction of loss reduction and then a line search is used to find the step length.
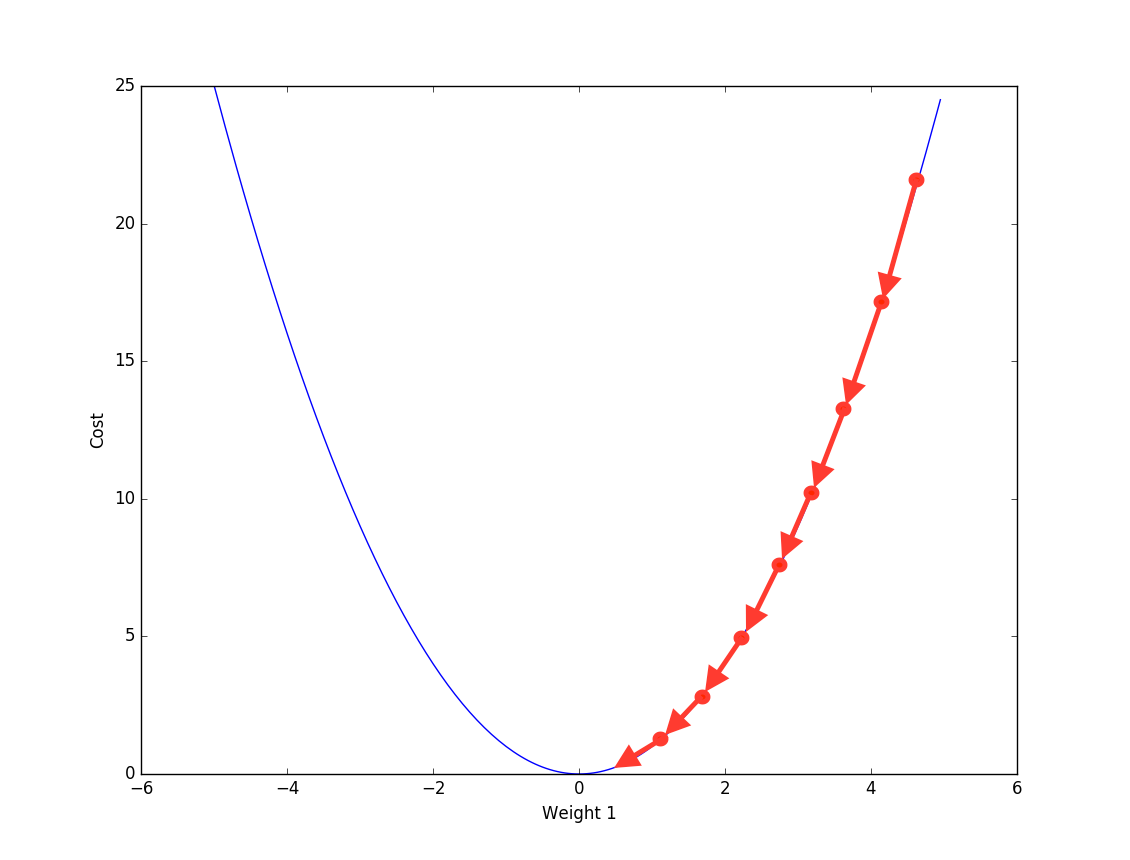
So far we have already mentioned 2 main components in Gradient Boosting machine:
- Boosting: \(F_m(x) = \sum \rho_m h(x;\alpha_m)\) final function is an additive model of multiple base learner.
- Optimization: Gradient descent is used as numeric optimization method.
There is one other important component, which we will cover later.
Gradient Boosting Basic
Here is the generic frame work of gradient Boosting algorithm.
Algorithm 1: Gradient_Boost
- \(F_0(x) = argmin \sum_i^N{L(y_i,p)}\)
- For m = 1 to M do :
A. $\hat{y_i} = - {[\frac{\partial{L(y_i, F(x_i)) }}{\partial{F(x_i ) } } ] } $ $ \text{ where \(F(x) = F_{m-1}(x)\)} $
B. \(a_m = argmin_{\alpha, \beta}\sum_i^N{[\hat{y_i} - \beta h(x_i;
\alpha)]^2}\)C. \(\rho_m = argmin_{\rho}\sum_i^N{L(y_i, F_{m-1}(x_i) + \rho h(x_i; \alpha_m))}\)
D. $ F_m(x_i) = F_{m-1}(x_i) + \rho h(x_i; \alpha_m))$
- \(F_m(x_i)\) will be final prediction
Let's go through all the above steps one by one.
Step1 we initialize the additive model, usually we can initialize with 0.
Then at each iteration:
A. Calculate negative gradient of current additive function
B. Fit a base learner to approximate negative gradient (direction of loss reduction).
C. find optimal coefficient for above base learner (step length), by minimizing the loss function.
D. update the additive function with new base learner and coefficient.
Step B is very important in the algorithm. Because given \(x\) and current additive function \(F_{m-1}(x)\), for each \(x_i\) we will have a empirical gradient. However we don't want the model to over-fit the training data. That's why an approximation estimation is used. Above we use Least-squares-Loss to fit the base leaner to the negative gradient, because least-squares estimates conditional expectation - \(E(\hat{y}|x)\)
Next let's use 2 loss function as example:
Least-Squares, all steps follow above algorithm, and we can further specify 2(A) 2(B) 2(C) as following:
Algorithm 2: LS_Boost
- \(F_0(x) = argmin \sum_i^N{L(y_i,p)}\)
- For m = 1 to M do :
A. $\hat{y_i} = y_i - F(x_i) \text{ where \(F(x) = F_{m-1}(x)\)}$
B. \(a_m = argmin_{\alpha, \beta}\sum_i^N{[\hat{y_i} - \beta h(x_i;\alpha)]^2}\)
C. $\rho_m =\beta $
D. $ F_m(x_i) = F_{m-1}(x_i) + \rho h(x_i; \alpha_m))$
Here negative gradient is simply the current residual. Since we already use least-square regression to fit base learner to the residual, where we not only get the base learner but also the coefficient. Therefore step 2(c) is no longer needed.
Least-absolute-deviation, all steps follow above algorithm, and we can further specify 2(A) 2(B) 2(C) as following:
Algorithm 3: LAD_Boost
- \(F_0(x) = argmin \sum_i^N{L(y_i,p)}\)
- For m = 1 to M do :
A. $\hat{y_i} = sign( y_i - F(x_i) ) \text{ where \(F(x) = F_{m-1}(x)\)}$
B. \(a_m = argmin_{\alpha, \beta}\sum_i^N{[\hat{y_i} - \beta h(x_i;\alpha)]^2}\)
C.\(\rho_m = argmin_{\rho}\sum_i^N{ |\hat{y_i} - \rho h(x_i; \alpha_m)|}\)
D. $ F_m(x_i) = F_{m-1}(x_i) + \rho h(x_i; \alpha_m))$
Here negative gradient is the sign of residual. And as before we fit base learner to the sign via least-squares regression.
Boosting married Tree
Now is time to reveal the last component of Gradient Boosting Machine - using Regression Tree as base learner. Same as AdaBoost, Gradient Boosting have more attractive features when it uses regression tree as base learner.
Therefore we can further represent each base leaner as an additive model (we mentioned in the previous Decision Tree Post) like below
\]
Instead of using linear regression, we fit a regression tree against negative gradient. With Least-square as loss function, \(b_j\) will the be average of gradient in each leaf.
We can further simplify this by combining \(b_j\) with coefficient \(\rho_m\), as following:
F_m(x_i) &= F_{m-1}(x_i) + \rho\sum_{j=1}^J{b_{jm} I(x \in R_{jm})}\\
F_m(x_i) &= F_{m-1}(x_i) + \sum_{j=1}^J{\lambda_{jm} I(x \in R_{jm})}
\end{align}
\]
The above transformation shows that when we fit the regression tree, we only need the node split, not the leaf assignment. In other words we are fitting Unit Gradient.
Later given the sample in each leaf, we calculate the leaf assignment by minimizing the loss function within each leaf. This will return the same result as minimizing the loss function over all sample, because all leaves are disjoint.
\]
This is also why each base learner (tree) needs to be a weak leaner. If the tree is deep enough that each leaf has only 1 sample, then we are just calculating empirical gradient of the training data (over-fitting).
From here we will use tree as default base learner, and let's go through all kinds of loss functions supported by Sklearn.
Sklean source code - loss function
Sklearn supports 7 loss function in total, 3 for classification and 4 for regression, see below
| Type | Loss | Estimator |
|---|---|---|
| ClassificationLossFunction | BinomialDeviance | LogOddsEstimator |
| ClassificationLossFunction | MultinomialDeviance | PriorProbabilityEstimator |
| ClassificationLossFunction | MultinomialDeviance | PriorProbabilityEstimator |
| RegressionLossFunction | LeastSquaresError | MeanEstimator |
| RegressionLossFunction | LeastAbsoluteError | QuantileEstimator |
| RegressionLossFunction | HuberLossFunction | QuantileEstimator |
| RegressionLossFunction | QuantileLossFunction | QuantileEstimator |
Here the Estimator calculates the prediction given training sample, including average, quantile(median), probability of each classes, or log-odds (\(\log p/1-p\)).
Each loss classes supports several methods, including calculating loss, negative gradient and update terminal regions.
For update_terminal_regions method, it first assign each sample to a leaf, perform line search to get leaf assignment and then update the prediction accordingly, see below:
class LossFunction(six.with_metaclass(ABCMeta, object)):
def update_terminal_regions(self, tree, X, y, residual, y_pred,
sample_weight, sample_mask,
learning_rate=1.0, k=0):
terminal_regions = tree.apply(X)
for leaf in np.where(tree.children_left == TREE_LEAF)[0]:
self._update_terminal_region(tree, masked_terminal_regions,
leaf, X, y, residual,
y_pred[:, k], sample_weight)
y_pred[:, k] += (learning_rate
* tree.value[:, 0, 0].take(terminal_regions, axis=0))
Next let's take a deeper dive into all loss functions.
Regression
1. Least-squares (LS)
Algorithm 4: LS_TreeBoost
- \(F_0(x) = mean(y_i)\)
- For m = 1 to M do :
A. $\hat{y_i} = y_i - F(x_i) \text{ where \(F(x) = F_{m-1}(x)\)}$
B. \(\{R_{jm}\}_1^J = \text{J terminal node tree}(\{\hat{y_i}, x_i\}_1^N)\)
C. $\lambda_{jm} = mean_{x_i \in R_{jm}}{ y_i - F(x_i)} $
D. $ F_m(x_i) = F_{m-1}(x_i) + \sum_{j=1}^J \lambda_{jm} I(x_i \in R_{jm})$
Compare algorithm 4 with algorithm 2, the difference lies in how the negative gradient is estimated, linear regression vs. regression tree. And how the coefficient is calculated, regression coefficient vs. sample mean in each leaf. And one attractive of LS is that it can get leaf assignment directly from gradient approximation. Therefore in the class update_terminal_regions is directly called to update prediction.
class LeastSquaresError(RegressionLossFunction):
def init_estimator(self):
return MeanEstimator()
def __call__(self, y, pred, sample_weight=None):
return np.mean((y - pred.ravel()) ** 2.0)
def negative_gradient(self, y, pred, **kargs):
return y - pred.ravel()
def update_terminal_regions(self, tree, X, y, residual, y_pred, learning_rate=1.0, k=0):
y_pred[:, k] += learning_rate * tree.predict(X).ravel()
2. Least-absolute-deviation (LAD)
Algorithm 5: LAD_TreeBoost
- \(F_0(x) = median(y_i)\)
- For m = 1 to M do :
A. $\hat{y_i} = sign( y_i - F(x_i) ) \text{ where \(F(x) = F_{m-1}(x)\)}$
B. \(\{R_{jm}\}_1^J = \text{J terminal node tree}(\{\hat{y_i}, x_i\}_1^N)\)
C. $\lambda_{jm} = median_{x_i \in R_{jm}}{ y_i - F(x_i)} $
D. $ F_m(x_i) = F_{m-1}(x_i) + \sum_{j=1}^J \lambda_{jm} I(x_i \in R_{jm})$
From algorithm implementation, we can see LAD is a very robust algorithm, which only use sample order to adjust the prediction accordingly. In _update_terminal_region, sample median in each leaf are used as leaf assignment.
class LeastAbsoluteError(RegressionLossFunction):
def init_estimator(self):
return QuantileEstimator(alpha=0.5)
def __call__(self, y, pred, sample_weight=None):
return np.abs(y - pred.ravel()).mean()
def negative_gradient(self, y, pred, **kargs):
pred = pred.ravel()
return 2.0 * (y - pred > 0.0) - 1.0
def _update_terminal_region(self, tree, terminal_regions, leaf, X, y, residual, pred, sample_weight):
terminal_region = np.where(terminal_regions == leaf)[0]
sample_weight = sample_weight.take(terminal_region, axis=0)
diff = y.take(terminal_region, axis=0) - pred.take(terminal_region, axis=0)
tree.value[leaf, 0, 0] = _weighted_percentile(diff, sample_weight, percentile=50)
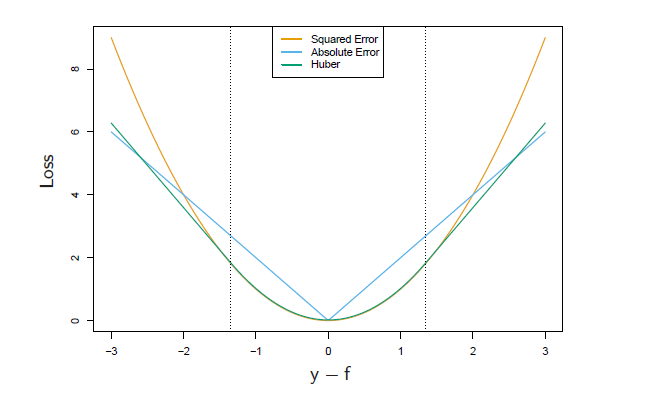
3. Huber Loss (M-Regression)
Huber is between LS and LAD, which uses LS when the error is smaller than certain quantile and use LAD when the error exceeds certain quantile, see below:
L(y,F) =
\begin{cases}
\frac{1}{2}(y-F)^2 & \quad & |y-F| \leq \delta \\
\delta(|y-F| - \delta/2) & \quad & |y-F| > \delta
\end{cases}
\end{align}
\]
where \(\delta\) is usually a certain quantile of absolute error. In additive model, we will use current additive prediction to estimate this quantile.
\]
Then we can further calculate negative gradient, and use a regression tree to approximate it.
\tilde{y} =\begin{cases}
y-F \quad & |y-F| \leq \delta \\
\delta \cdot sign(y-F) \quad & |y-F| > \delta
\end{cases}
\end{align}
\]
In the end we need to minimize loss in each leaf, which can be a little tricky with Huber loss function. For LAD, optimal leaf value is the median, for LS, optimal leaf value is the mean. For Huber an approximation method is used, where we use the sample within \(\delta\) to adjust the overall median, like following:
\]
where \(\tilde{\gamma_{jm}} = median(\gamma_{jm}) = median(y-F)\) is the LAD estimation - median of residual in each leaf.
Algorithm 6: M_TreeBoost
- \(F_0(x) = median(y_i)\)
- For m = 1 to M do :
A. \(r_{m-1} = y_i - F(x_i)\)
B. \(\delta_m = quantile_{\alpha}(|r_{m-1}|_1^N)\)
C. $
\tilde{y} =\begin{cases}
y-F \quad & |y-F| \leq \delta_m \
\delta_m \cdot sign(y-F) \quad & |y-F| > \delta_m
\end{cases} $D. \(\{R_{jm}\}_1^J = \text{J terminal node tree}(\{\tilde{y}, x_i\}_1^N)\)
E. \(\lambda_{jm}= \tilde{\gamma_{jm}} + \frac{1}{N_{jm}}\sum_{x \in R_{jm}} sign(\gamma_{m-1}(x_i) -\tilde{\gamma_{jm}} ) \cdot min(\delta_m, abs(\gamma_{m-1}(x_i) -\tilde{\gamma_{jm}}))\)
F. $ F_m(x_i) = F_{m-1}(x_i) + \sum_{j=1}^J \lambda_{jm} I(x_i \in R_{jm})$
In application, I found Huber-Loss to be a very powerful loss function. Because it is not that sensitive to the outlier, but is able to capture more information than just sample order as LAD.
class HuberLossFunction(RegressionLossFunction):
def init_estimator(self):
return QuantileEstimator(alpha=0.5)
def __call__(self, y, pred, sample_weight=None):
pred = pred.ravel()
diff = y - pred
gamma = stats.scoreatpercentile(np.abs(diff), self.alpha * 100)
gamma_mask = np.abs(diff) <= gamma
sq_loss = np.sum(0.5 * diff[gamma_mask] ** 2.0)
lin_loss = np.sum(gamma * (np.abs(diff[~gamma_mask]) - gamma / 2.0))
loss = (sq_loss + lin_loss) / y.shape[0]
return loss
def negative_gradient(self, y, pred, sample_weight=None, **kargs):
pred = pred.ravel()
diff = y - pred
gamma = stats.scoreatpercentile(np.abs(diff), self.alpha * 100)
gamma_mask = np.abs(diff) <= gamma
residual = np.zeros((y.shape[0],), dtype=np.float64)
residual[gamma_mask] = diff[gamma_mask]
residual[~gamma_mask] = gamma * np.sign(diff[~gamma_mask])
self.gamma = gamma
return residual
def _update_terminal_region(self, tree, terminal_regions, leaf, X, y,
residual, pred, sample_weight):
terminal_region = np.where(terminal_regions == leaf)[0]
sample_weight = sample_weight.take(terminal_region, axis=0)
gamma = self.gamma
diff = (y.take(terminal_region, axis=0)
- pred.take(terminal_region, axis=0))
median = _weighted_percentile(diff, sample_weight, percentile=50)
diff_minus_median = diff - median
tree.value[leaf, 0] = median + np.mean(
np.sign(diff_minus_median) *
np.minimum(np.abs(diff_minus_median), gamma))
Classification
1. 2-class classification
For binomial classification, our calculation below will be slightly different from Freud's paper. Because Sklearn use \(y \in \{0,1\}\), while in Freud's paper \(y \in \{-1,1\}\) is used. Of course, whichever you use, you should get the same result. You need to make sure that your calculation is consistent. Relevant bug was spotted in Sklearn before.
Do you still recall how is the binomial log-likelihood defined? We define current prediction as log-odds.
F & = \log(\frac{P(y=1|x)}{P(y=0|x)}) \\
P &= \frac{1}{1+e^{-F}} \\
\end{align}
\]
And use above to calculate negative log-likelihood function, we get following
L(y,F) & = - E(y\log{p} + (1-y)log{(1-p)} ) \\
& = - E( y\log{\frac{p}{1-p}} + \log({1-p}) ) \\
& = - E( yF - \log{(1+e^F)} )
\end{align}
\]
The negative gradient is below, I also give a general version, which is in line with later k-class regression
\tilde{y} & = y - \frac{1}{1+ e^{-,F}} = y - \sigma(F) = y - p
\end{align}
\]
In the end we calculate leaf assignment by minimizing the loss in each leaf
\]
There is no close solution to above function, a second-order Newton Raphson is used to approximate
Quick Note - Newton Raphson
\(f(x + \epsilon) \approx f(x) + f'(x)\epsilon + \frac{1}{2}f''(x)\epsilon^2\) 2nd order Taylor expansion
To get optimal value, \(f'(x) = 0\)
we will get \(\epsilon = -\frac{f'(x)}{f''(x)}\)
\lambda_{jm} & = \sum_{x \in R_{jm}}(y - \frac{1}{1+ e^{-F}} )/(\frac{1}{1+ e^{-F}} \cdot \frac{1}{1+ e^{F}}) \\
& = \sum_{x \in R_{jm}} \tilde{y} / ( (y-\tilde{y}) \cdot (1-y+\tilde{y}) )
\end{align}
\]
Algorithm 7: L2_TreeBoost
- \(F_0(x) = 0\)
- For m = 1 to M do :
A. $\tilde{y} = y - \frac{1}{1+ e^{-F}} \text{ where \(F(x) = F_{m-1}(x)\)}$
B. \(\{R_{jm}\}_1^J = \text{J terminal node tree}(\{\hat{y_i}, x_i\}_1^N)\)
C. $\lambda_{jm} = \sum_{x \in R_{jm}} \tilde{y} / ( (y-\tilde{y}) \cdot (1-y+\tilde{y})) $
D. $ F_m(x_i) = F_{m-1}(x_i) + \sum_{j=1}^J \lambda_{jm} I(x_i \in R_{jm})$
class BinomialDeviance(ClassificationLossFunction):
def init_estimator(self):
return LogOddsEstimator()
def __call__(self, y, pred, sample_weight=None):
pred = pred.ravel()
return -2.0 * np.mean((y * pred) - np.logaddexp(0.0, pred)) #logaddexp(0, v) == log(1.0 + exp(v))
def negative_gradient(self, y, pred, **kargs):
return y - expit(pred.ravel()) # sigmoid function
def _update_terminal_region(self, tree, terminal_regions, leaf, X, y,
residual, pred, sample_weight):
terminal_region = np.where(terminal_regions == leaf)[0]
residual = residual.take(terminal_region, axis=0)
y = y.take(terminal_region, axis=0)
sample_weight = sample_weight.take(terminal_region, axis=0)
numerator = np.sum(sample_weight * residual)
denominator = np.sum(sample_weight * (y - residual) * (1 - y + residual))
tree.value[leaf, 0, 0] = numerator / denominator
2. k-class classification
Loss function is defined in the same ways with multiclass:
\]
where \(p_k(x) = exp(F_k(x))/\sum_{l=1}^Kexp(F_l(x))\)
Therefore we will get the loss function nad negative gradient as following:
L(y,F) &= -\sum_{k=1}^K y_kF_k(x) - log(\sum_{l=1}^Kexp(F_l(x)))\\
\tilde{y_i} &= y_{ki} - p_{k,m-1}(x_i)
\end{align}
\]
Algorithm 8: LK_TreeBoost
- \(F_0(x) = 0\)
- For m = 1 to M do :
A. \(p_k(x) = exp(F_k(x))/\sum_{l=1}^Kexp(F_l(x))\)
B. \(\tilde{y_i} = y_i - p_k(x_i)\)
B. \(\{R_{jm}\}_1^J = \text{J terminal node tree}(\{\hat{y_i}, x_i\}_1^N)\)
C. $\lambda_{jm} = \sum_{x \in R_{jm}} \tilde{y} / ( (y-\tilde{y}) \cdot (1-y+\tilde{y})) $
D. $ F_m(x_i) = F_{m-1}(x_i) + \sum_{j=1}^J \lambda_{jm} I(x_i \in R_{jm})$
sklearn source code - GBM Framework
Base Learner
Base learner in each iteration is trained via fit_stage method. A Decision Tree is trained to approximate negative gradient given current additive function. And then leaf assignment is calculated to minimize loss function in each leaf.
def _fit_stage(self, i, X, y, y_pred, sample_weight, sample_mask, random_state, X_idx_sorted, X_csc=None, X_csr=None):
assert sample_mask.dtype == np.bool
loss = self.loss_
original_y = y
for k in range(loss.K):
residual = loss.negative_gradient(y, y_pred, k=k, sample_weight=sample_weight)
tree = DecisionTreeRegressor(
criterion=self.criterion,
splitter='best',
max_depth=self.max_depth,
min_samples_split=self.min_samples_split,
min_samples_leaf=self.min_samples_leaf,
min_weight_fraction_leaf=self.min_weight_fraction_leaf,
min_impurity_decrease=self.min_impurity_decrease,
min_impurity_split=self.min_impurity_split,
max_features=self.max_features,
max_leaf_nodes=self.max_leaf_nodes,
random_state=random_state,
presort=self.presort)
tree.fit(X, residual, sample_weight=sample_weight,
check_input=False, X_idx_sorted=X_idx_sorted)
loss.update_terminal_regions(tree.tree_, X, y, residual, y_pred,sample_weight, sample_mask, self.learning_rate, k=k)
self.estimators_[i, k] = tree
return y_pred
Boosting
Boosting is performed upon above base learner via fit_stages method. In each iteration a new base learner is trained given current additive function.
n_estimators specifies the number of iteration (# of base learner).
subsample indicates the split between training sample and validation sample. Base learner is trained against training sample, and we use untouched validation sample for performance.
def _fit_stages(self, X, y, y_pred, sample_weight, random_state,
begin_at_stage=0, monitor=None, X_idx_sorted=None):
n_samples = X.shape[0]
do_oob = self.subsample < 1.0
sample_mask = np.ones((n_samples, ), dtype=np.bool)
n_inbag = max(1, int(self.subsample * n_samples))
loss_ = self.loss_
i = begin_at_stage
for i in range(begin_at_stage, self.n_estimators):
if do_oob:
sample_mask = _random_sample_mask(n_samples, n_inbag,
random_state)
old_oob_score = loss_(y[~sample_mask],
y_pred[~sample_mask],
sample_weight[~sample_mask])
y_pred = self._fit_stage(i, X, y, y_pred, sample_weight, sample_mask, random_state, X_idx_sorted, X_csc, X_csr)
if do_oob:
self.train_score_[i] = loss_(y[sample_mask],
y_pred[sample_mask], sample_weight[sample_mask])
self.oob_improvement_[i] = (
old_oob_score - loss_(y[~sample_mask],
y_pred[~sample_mask], sample_weight[~sample_mask]))
return i + 1
Reference
- J. Friedman, Greedy Function Approximation: A Gradient Boosting Machine, The Annals of Statistics, Vol. 29, No. 5, 2001.
- J. Friedman, Stochastic Gradient Boosting, 1999
- T. Hastie, R. Tibshirani and J. Friedman. Elements of Statistical Learning Ed. 2, Springer, 2009.
- Bishop, Pattern Recognition and Machine Learning 2006
- scikit-learn tutorial http://scikit-learn.org/stable/modules/generated/sklearn.ensemble.GradientBoostingClassifier.html
Tree - Gradient Boosting Machine with sklearn source code的更多相关文章
- Tree - AdaBoost with sklearn source code
In the previous post we addressed some issue of decision tree, including instability, lack of smooth ...
- Tree - Decision Tree with sklearn source code
After talking about Information theory, now let's come to one of its application - Decision Tree! No ...
- 论文笔记:GREEDY FUNCTION APPROXIMATION: A GRADIENT BOOSTING MACHINE
Boost是集成学习方法中的代表思想之一,核心的思想是不断的迭代.boost通常采用改变训练数据的概率分布,针对不同的训练数据分布调用弱学习算法学习一组弱分类器.在多次迭代的过程中,当前次迭代所用的训 ...
- Python中Gradient Boosting Machine(GBM)调参方法详解
原文地址:Complete Guide to Parameter Tuning in Gradient Boosting (GBM) in Python by Aarshay Jain 原文翻译与校对 ...
- 机器学习--Gradient Boosting Machine(GBM)调参方法详解
一.GBM参数 总的来说GBM的参数可以被归为三类: 树参数:调节模型中每个决策树的性质 Boosting参数:调节模型中boosting的操作 其他模型参数:调节模型总体的各项运作 1.树参数 现在 ...
- Greedy Function Approximation:A Gradient Boosting Machine
https://statweb.stanford.edu/~jhf/ftp/trebst.pdf page10 90% to 95% of the observations were often de ...
- A Gentle Introduction to the Gradient Boosting Algorithm for Machine Learning
A Gentle Introduction to the Gradient Boosting Algorithm for Machine Learning by Jason Brownlee on S ...
- Gradient Boosting Decision Tree学习
Gradient Boosting Decision Tree,即梯度提升树,简称GBDT,也叫GBRT(Gradient Boosting Regression Tree),也称为Multiple ...
- How to Configure the Gradient Boosting Algorithm
How to Configure the Gradient Boosting Algorithm by Jason Brownlee on September 12, 2016 in XGBoost ...
随机推荐
- (Les17 移动数据)expdp/impdp
oracle 11.2.0 expdp/impdp 数据泵参数 expdp参数=========================================================== ...
- Python 基础 函数
python 什么是函数 Python不但能非常灵活地定义函数,而且本身内置了很多有用的函数,可以直接调用. python 函数的调用 Python内置了很多有用的函数,我们可以直接调用. 要调用 ...
- 页面缓存优化系列一(expires,cache-control 解读)
参考文章:https://developer.mozilla.org/zh-CN/docs/Web/HTTP/Headers/Cache-Control http://caibaojian.com/s ...
- jquery中的 append , after , prepend , before 区别
jQuery append() 方法在被选元素的结尾插入内容. jQuery prepend() 方法在被选元素的开头插入内容. jQuery after() 方法在被选元素之后插入内容. jQuer ...
- MySQL数据库实验:任务二 表数据的插入、修改及删除
目录 任务二 表数据的插入.修改及删除 一.利用界面工具插入数据 二.数据更新 (一)利用MySQL命令行窗口更新数据 (二)利用Navicat for MySQL客户端工具更新数据 三.数据库的备份 ...
- 使用cmd时cd命令失效
使用cmd时cd命令失效 近日使用cmd时总是出现无法cd到指定目录的情况 如下图所示 输入cd命令后依旧停留在原始路径 解决方法: 输入 cd D:\CE-5\Training_Sanple\n ...
- My First
刚入职不到2个月吧,还在实习,月底拿毕业证转正.工作期间遇到很多麻烦问题,有的解决了,有的解决不了,换了个方法实现,挺无奈的.弄个博客,记录下平常遇到的问题和解决方式,也省的每次拿个笔记下来了…… 公 ...
- 20155202 《Java程序设计》实验二(面向对象程序设计)实验报告
20155202 <Java程序设计>实验二(面向对象程序设计)实验报告 代码托管 实验内容 初步掌握单元测试和TDD 理解并掌握面向对象三要素:封装.继承.多态 初步掌握UML建模 熟悉 ...
- C语言字节对齐问题详解(对齐、字节序、网络序等)
首先说明一下,本文是转载自: http://www.cnblogs.com/clover-toeic/p/3853132.html 博客园用的少,不知道怎么发布转载文章,只能暂时这样了. 引言 考虑下 ...
- 《JAVA程序设计》 实验二 Java面向对象程序设计
<JAVA程序设计> 实验二 Java面向对象程序设计 实验内容 初步掌握单元测试和TDD 理解并掌握面向对象三要素:封装.继承.多态 初步掌握UML建模 熟悉S.O.L.I.D原则 了解 ...
