DNA sequence HDU - 1560
DNA sequence
Time Limit: 15000/5000 MS (Java/Others) Memory Limit: 32768/32768 K (Java/Others)
Total Submission(s): 4217 Accepted Submission(s): 2020
For example, given "ACGT","ATGC","CGTT" and "CAGT", you can make a sequence in the following way. It is the shortest but may be not the only one.
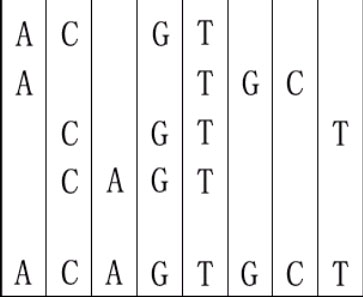
4
ACGT
ATGC
CGTT
CAGT
#include<iostream>
#include<cstdio>
#include<cstdlib>
#include<sstream>
#include<cstring>
#include<string>
#include<vector>
#include<set>
#include<stack>
#include<queue>
#include<map>
#include<cmath>
#include<algorithm>
using namespace std;
#define inf 0x3f3f3f3f
#define ll long long
#define MAX_N 1000005
#define gcd(a,b) __gcd(a,b)
#define mem(a,x) memset(a,x,sizeof(a))
#define mid(a,b) a+b/2
#define stol(a) atoi(a.c_str())//string to long
string temp[];
string c = "ACGT";
int pos[];
int len[];
int deep;
int n;
int ans(){
int maxn = ;
for(int i = ; i < n; i++){
maxn = max(maxn,len[i]-pos[i]);
}
return maxn;
}
int dfs(int step){
if(step + ans() > deep)
return ;
if(!ans())//满足构造序列包含所有序列
return ;
int temp1[];
for(int i = ; i < ; i++){ int flag = ;
for(int j = ; j < n; j++)
temp1[j] = pos[j]; for(int j = ; j < n; j++){
if(temp[j][pos[j]] == c[i]){
flag = ;
pos[j]++;
}
}
if(flag){
if(dfs(step+))
return ;
for(int j = ; j < n; j++)
pos[j] = temp1[j];
}
}
return ;
}
int main(){
// freopen("D:\\in.txt","r",stdin);
int T;
scanf("%d",&T);
while(T--){
scanf("%d",&n);
deep = ;
for(int i = ; i < n; i++){
cin>>temp[i];
len[i] = temp[i].length();
pos[i] = ;
deep = max(deep,len[i]);
}
while(){
if(dfs())
break;
++deep;
}
printf("%d\n",deep);
}
return ;
}
DNA sequence HDU - 1560的更多相关文章
- DNA sequence HDU - 1560(IDA*,迭代加深搜索)
题目大意:有n个DNA序列,构造一个新的序列,使得这n个DNA序列都是它的子序列,然后输出最小长度. 题解:第一次接触IDA*算法,感觉~~好暴力!!思路:维护一个数组pos[i],表示第i个串该匹配 ...
- IDA*、剪枝、较难搜索、扫描——DNA sequence HDU - 1560
万恶之源 翻译 题意就是给出N个DNA序列,要求出一个包含这n个序列的最短序列是多长 这是一道搜索题,为什么呢?从样例可以感受到,我们应该从左往右"扫描",从n个DNA序列中取出某 ...
- G - DNA sequence HDU - 1560
题目链接: https://vjudge.net/contest/254151#problem/G AC代码: #include<iostream> #include<cstring ...
- hdu 1560 DNA sequence(搜索)
http://acm.hdu.edu.cn/showproblem.php?pid=1560 DNA sequence Time Limit: 15000/5000 MS (Java/Others) ...
- HDU 1560 DNA sequence(DNA序列)
HDU 1560 DNA sequence(DNA序列) Time Limit: 15000/5000 MS (Java/Others) Memory Limit: 32768/32768 K ...
- hdu 1560 DNA sequence(迭代加深搜索)
DNA sequence Time Limit : 15000/5000ms (Java/Other) Memory Limit : 32768/32768K (Java/Other) Total ...
- 【HDU - 1560】DNA sequence (dfs+回溯)
DNA sequence 直接中文了 题目描述 21世纪是生物科技飞速发展的时代.我们都知道基因是由DNA组成的,而DNA的基本组成单位是A,C,G,T.在现代生物分子计算中,如何找到DNA之间的最长 ...
- HDU1560 DNA sequence —— IDA*算法
题目链接:http://acm.hdu.edu.cn/showproblem.php?pid=1560 DNA sequence Time Limit: 15000/5000 MS (Java/Oth ...
- POJ 2778 DNA Sequence(AC自动机+矩阵加速)
DNA Sequence Time Limit: 1000MS Memory Limit: 65536K Total Submissions: 9899 Accepted: 3717 Desc ...
随机推荐
- 401认证钓鱼demo
<?php //@b4dboy if(!isset($_SERVER['PHP_AUTH_USER']) && !isset($_SERVER['PHP_AUTH_PW'])) ...
- ununtu 16.04 下的 VsCode 下载与安装
Vscode发现用包下载显示找不到网页,于是只有继续折腾. 折腾如下: ubuntu-desktop You can update your system with unsupported packa ...
- F5 BIG-IP之一前期随笔(应用交付网络产品)
ADN:应用交付网络 TMOS: Traffic managment operation system (流量管理系统) 一个实时的全应用代理流量管理操作系统 PVA ...
- 浙江省第十六届大学生ACM程序设计竞赛部分题解
E .Sequence in the Pocket sol:将数组copy一份,然后sort一下,找寻后面最多多少个元素在原数组中保持有序,用总个数减去已经有序的就是我们需要移动的次数. 思维题 #i ...
- cs231n spring 2017 lecture3 Loss Functions and Optimization
1. Loss function是用来量化评估当前预测的好坏,loss function越小表明预测越好. 几种典型的loss function: 1)Multiclass SVM loss:一般的S ...
- [LC] 367. Valid Perfect Square
Given a positive integer num, write a function which returns True if num is a perfect square else Fa ...
- linux debain systemd 开机启动 nodejs 兼容原initd启动 forever 开机自启
布署环境为debian 7.5 布署到一台新机器,系统版本为debian 8.0 原启动项 /etc/init.d/mongo_service 开机居然无法自起 开机自启动不能用了,看页面输出就发觉不 ...
- HDU-2802-F(N)
看到这题讨论版里有说用公式的有说用循环节的,但是个人觉得这两种方法都不靠谱,比赛场上做这种题能直接推出公式需要很强数学功底,而循环节的方法如果循环节比较大就不太好发现了.这种已知通项公式的题还是用矩阵 ...
- <SCOI2008>奖励关
emmm第一道期望dp+个状压 真有趣.. #include<cstdio> #include<iostream> #include<cstring> #inclu ...
- configure: error: C compiler cannot create executables报错处理
在测试环境安装php的imagick扩展在执行./configure生成编译文件时出现报错如下: 通过查看config.log发现有报错,在网上经验教程里发现前面的报错不管,直奔最后的报错即可,发现是 ...
