Human Gene Functions POJ 1080 最长公共子序列变形
Description
A human gene can be identified through a series of time-consuming biological experiments, often with the help of computer programs. Once a sequence of a gene is obtained, the next job is to determine its function.
One of the methods for biologists to use in determining the function of a new gene sequence that they have just identified is to search a database with the new gene as a query. The database to be searched stores many gene sequences and their functions – many researchers have been submitting their genes and functions to the database and the database is freely accessible through the Internet.
A database search will return a list of gene sequences from the database that are similar to the query gene.
Biologists assume that sequence similarity often implies functional similarity. So, the function of the new gene might be one of the functions that the genes from the list have. To exactly determine which one is the right one another series of biological experiments will be needed.
Your job is to make a program that compares two genes and determines their similarity as explained below. Your program may be used as a part of the database search if you can provide an efficient one.
Given two genes AGTGATG and GTTAG, how similar are they? One of the methods to measure the similarity
of two genes is called alignment. In an alignment, spaces are inserted, if necessary, in appropriate positions of
the genes to make them equally long and score the resulting genes according to a scoring matrix.
For example, one space is inserted into AGTGATG to result in AGTGAT-G, and three spaces are inserted into GTTAG to result in –GT--TAG. A space is denoted by a minus sign (-). The two genes are now of equal
length. These two strings are aligned:
AGTGAT-G
-GT--TAG
In this alignment, there are four matches, namely, G in the second position, T in the third, T in the sixth, and G in the eighth. Each pair of aligned characters is assigned a score according to the following scoring matrix. 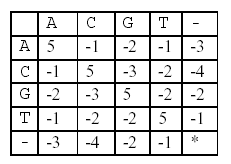
denotes that a space-space match is not allowed. The score of the alignment above is (-3)+5+5+(-2)+(-3)+5+(-3)+5=9.
Of course, many other alignments are possible. One is shown below (a different number of spaces are inserted into different positions):
AGTGATG
-GTTA-G
This alignment gives a score of (-3)+5+5+(-2)+5+(-1) +5=14. So, this one is better than the previous one. As a matter of fact, this one is optimal since no other alignment can have a higher score. So, it is said that the
similarity of the two genes is 14.
Input
Output
Sample Input
2
7 AGTGATG
5 GTTAG
7 AGCTATT
9 AGCTTTAAA
Sample Output
14
21
Source
注意边界条件的处理!
#include<iostream>
#include<cstdio>
#include<cmath>
#include<cstring>
#include<sstream>
#include<algorithm>
#include<queue>
#include<deque>
#include<iomanip>
#include<vector>
#include<cmath>
#include<map>
#include<stack>
#include<set>
#include<fstream>
#include<memory>
#include<list>
#include<string>
using namespace std;
typedef long long LL;
typedef unsigned long long ULL;
#define MAXN 109
#define MOD 1000000
#define INF 1000000009
#define eps 0.00000001
/*
最长子序列求出来
然后和原序列匹配
*/
char a[MAXN], b[MAXN];
int l1, l2, dp[MAXN][MAXN], g[MAXN][MAXN];
void Init()
{
g['A']['A'] = g['C']['C'] = g['T']['T'] = g['G']['G'] = ;
g['A']['C'] = g['C']['A'] = g['A']['T'] = g['T']['A'] = -;
g['A']['G'] = g['G']['A'] = -;
g['C']['T'] = g['T']['C'] = -;
g['G']['T'] = g['T']['G'] = -;
g['C']['G'] = g['G']['C'] = -;
g['A']['A' - ] = -;
g['C']['C' - ] = -;
g['G']['G' - ] = -;
g['T']['T' - ] = -;
}
int main()
{
Init();
int T; scanf("%d", &T);
while (T--)
{
memset(dp, - INF, sizeof(dp));
scanf("%d%s\n%d%s", &l1, a+, &l2, b+);
dp[][] = ;
for (int i = ; i <= l1; i++)
{
dp[i][] = dp[i - ][] + g[a[i]][a[i] - ];
}
for (int i = ; i <= l2; i++)
{
dp[][i] = dp[][i - ] + g[b[i]][b[i] - ];
}
for (int i = ; i <= l1; i++)
{
for (int j = ; j <= l2; j++)
{
if(a[i]==b[j])
dp[i][j] = dp[i - ][j - ] + g[a[i]][b[j]];
else
dp[i][j] = max(max(dp[i - ][j] + g[a[i]][a[i] - ], dp[i][j - ] + g[b[j]][b[j] - ]),dp[i-][j-]+g[a[i]][b[j]]);
}
}
//solve(l1, l2);
printf("%d\n", dp[l1][l2]);
}
}
Human Gene Functions POJ 1080 最长公共子序列变形的更多相关文章
- POJ 2250(最长公共子序列 变形)
Description In a few months the European Currency Union will become a reality. However, to join the ...
- POJ 1458 最长公共子序列(dp)
POJ 1458 最长公共子序列 题目大意:给出两个字符串,求出这样的一 个最长的公共子序列的长度:子序列 中的每个字符都能在两个原串中找到, 而且每个字符的先后顺序和原串中的 先后顺序一致. Sam ...
- poj 1080 Human Gene Functions (最长公共子序列变形)
题意:有两个代表基因序列的字符串s1和s2,在两个基因序列中通过添加"-"来使得两个序列等长:其中每对基因匹配时会形成题中图片所示匹配值,求所能得到的总的最大匹配值. 题解:这题运 ...
- POJ 1458 最长公共子序列
子序列就是子序列中的元素是母序列的子集,且子序列中元素的相对顺序和母序列相同. 题目要求便是寻找两个字符串的最长公共子序列. dp[i][j]表示字符串s1左i个字符和s2左j个字符的公共子序列的最大 ...
- POJ 1458 最长公共子序列 LCS
经典的最长公共子序列问题. 状态转移方程为 : if(x[i] == Y[j]) dp[i, j] = dp[i - 1, j - 1] +1 else dp[i, j] = max(dp[i - 1 ...
- 【简单dp】poj 1458 最长公共子序列【O(n^2)】【模板】
最长公共子序列可以用在下面的问题时:给你一个字符串,请问最少还需要添加多少个字符就可以让它编程一个回文串? 解法:ans=strlen(原串)-LCS(原串,反串); Sample Input abc ...
- hdu 1080 dp(最长公共子序列变形)
题意: 输入俩个字符串,怎样变换使其所有字符对和最大.(字符只有'A','C','G','T','-') 其中每对字符对应的值如下: 怎样配使和最大呢. 比如: A G T G A T G - G ...
- POJ 1159 Palindrome-最长公共子序列问题+滚动数组(dp数组的重复利用)(结合奇偶性)
Description A palindrome is a symmetrical string, that is, a string read identically from left to ri ...
- hdu1503 最长公共子序列变形
题目链接:http://acm.split.hdu.edu.cn/showproblem.php?pid=1503 题意:给出两个字符串 要求输出包含两个字符串的所有字母的最短序列.注意输出的顺序不能 ...
随机推荐
- Atlantis(坐标离散化)
http://poj.org/problem?id=1151 题意:给出矩形的左上角坐标和右下角坐标(坐标的y轴是向下的),求出矩形面积的并.. 今天好困啊..迷迷糊糊的听会神给讲了讲,敲完之后调试了 ...
- ARM VM安装Linux Diagnostic 2.3扩展
目前创建的Azure Linux虚拟机默认安装的是LAD 3.0,如果客户有特殊需求,可以通过如下方法安装LAD 2.3 1.在Azure Portal卸载LAD 3.0 2.使用Azure Powe ...
- JavaScript中的substr和Java中的substring
JavaScript::substr(index, length)从下标开始截取多少位,如果length为空,则截取到最后,-1倒数第一位,-2倒数第二位.... Java:substring(sta ...
- wcf 代理配置
<?xml version="1.0" encoding="utf-8"?> <configuration> <appSett ...
- Python 如何在csv中定位非数字和字母的符号
在数据清洗过程中,有时不仅希望去掉脏数据,更希望定位脏数据的位置,例如从csv里面定位非数字和字母单元格的位置,在使用isdigit().isalpha().isalnum()时无法判断浮点数,会将浮 ...
- 右边根据左边的高度自动居中只需要两行CSS就可以完成
右边根据左边的高度自动居中只需要两行CSS就可以完成 <style type="text/css" > div{ display: inline-block; vert ...
- Android 串口驱动和应用测试
这篇博客主要是通过一个简单的例子来了解Android的串口驱动和应用,为方便后续对Android串口服务和USB虚拟串口服务的了解.这个例子中,参考了<Linux Device Drivers& ...
- mongoDB 删除集合后,空间不释放的解决方法
mongoDB 删除集合后,空间不释放,添加新集合,没有重新利用之前删除集合所空出来的空间,也就是数据库大小只增不减. 方法有: 1.导出导入 dump & restore 2.修复数据库 r ...
- Hive扩展功能(六)--HPL/SQL(可使用存储过程)
软件环境: linux系统: CentOS6.7 Hadoop版本: 2.6.5 zookeeper版本: 3.4.8 主机配置: 一共m1, m2, m3这五部机, 每部主机的用户名都为centos ...
- (转)Hibernate的优化方案
http://blog.csdn.net/yerenyuan_pku/article/details/70768603 HQL优化 使用参数绑定 使用绑定参数的原因是让数据库一次解析SQL,对后续的 ...
