uva1368 DNA Consensus String
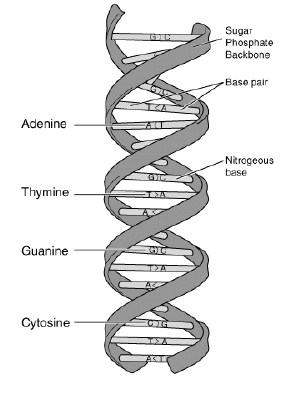 <tex2html_verbatim_mark> Figure 1.
<tex2html_verbatim_mark> Figure 1.DNA (Deoxyribonucleic Acid) is the molecule which contains the genetic instructions. It consists of four different nucleotides, namely Adenine, Thymine, Guanine, and Cytosine as shown in Figure 1. If we represent a nucleotide by its initial character, a DNA strand can be regarded as a long string (sequence of characters) consisting of the four characters A, T, G, and C. For example, assume we are given some part of a DNA strand which is composed of the following sequence of nucleotides:
``Thymine-Adenine-Adenine-Cytosine-Thymine-Guanine-Cytosine-Cytosine-Guanine-Adenine-Thymine"
Then we can represent the above DNA strand with the string ``TAACTGCCGAT." The biologist Prof. Ahn found that a gene X commonly exists in the DNA strands of five different kinds of animals, namely dogs, cats, horses, cows, and monkeys. He also discovered that the DNA sequences of the gene X from each animal were very alike. See Figure 2.
| DNA sequence of gene X | |
| Cat: | GCATATGGCTGTGCA |
| Dog: | GCAAATGGCTGTGCA |
| Horse: | GCTAATGGGTGTCCA |
| Cow: | GCAAATGGCTGTGCA |
| Monkey: | GCAAATCGGTGAGCA |
Prof. Ahn thought that humans might also have the gene X and decided to search for the DNA sequence of X in human DNA. However, before searching, he should define a representative DNA sequence of gene X because its sequences are not exactly the same in the DNA of the five animals. He decided to use the Hamming distance to define the representative sequence. The Hamming distance is the number of different characters at each position from two strings of equal length. For example, assume we are given the two strings ``AGCAT" and ``GGAAT." The Hamming distance of these two strings is 2 because the 1st and the 3rd characters of the two strings are different. Using the Hamming distance, we can define a representative string for a set of multiple strings of equal length. Given a set of strings S = s1,...,sm<tex2html_verbatim_mark> of length n<tex2html_verbatim_mark> , the consensus error between a string y<tex2html_verbatim_mark> of length n<tex2html_verbatim_mark> and the set S<tex2html_verbatim_mark> is the sum of the Hamming distances between y<tex2html_verbatim_mark> and eachsi<tex2html_verbatim_mark> in S<tex2html_verbatim_mark> . If the consensus error between y<tex2html_verbatim_mark> and S<tex2html_verbatim_mark> is the minimum among all possible strings y<tex2html_verbatim_mark> of length n<tex2html_verbatim_mark> , y<tex2html_verbatim_mark> is called a consensus string ofS<tex2html_verbatim_mark> . For example, given the three strings `` AGCAT" `` AGACT" and `` GGAAT" the consensus string of the given strings is `` AGAAT" because the sum of the Hamming distances between `` AGAAT" and the three strings is 3 which is minimal. (In this case, the consensus string is unique, but in general, there can be more than one consensus string.) We use the consensus string as a representative of the DNA sequence. For the example of Figure 2 above, a consensus string of gene X is `` GCAAATGGCTGTGCA" and the consensus error is 7.
Input
Your program is to read from standard input. The input consists of T<tex2html_verbatim_mark> test cases. The number of test cases T<tex2html_verbatim_mark> is given in the first line of the input. Each test case starts with a line containing two integers m<tex2html_verbatim_mark> and n<tex2html_verbatim_mark> which are separated by a single space. The integer m<tex2html_verbatim_mark>(4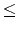 m
m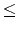 50)<tex2html_verbatim_mark>represents the number of DNA sequences and n<tex2html_verbatim_mark>(4
50)<tex2html_verbatim_mark>represents the number of DNA sequences and n<tex2html_verbatim_mark>(4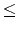 n
n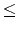 1000)<tex2html_verbatim_mark> represents the length of the DNA sequences, respectively. In each of the next m<tex2html_verbatim_mark> lines, each DNA sequence is given.
1000)<tex2html_verbatim_mark> represents the length of the DNA sequences, respectively. In each of the next m<tex2html_verbatim_mark> lines, each DNA sequence is given.
Output
Your program is to write to standard output. Print the consensus string in the first line of each case and the consensus error in the second line of each case. If there exists more than one consensus string, print the lexicographically smallest consensus string. The following shows sample input and output for three test cases.
Sample Input
3
5 8
TATGATAC
TAAGCTAC
AAAGATCC
TGAGATAC
TAAGATGT
4 10
ACGTACGTAC
CCGTACGTAG
GCGTACGTAT
TCGTACGTAA
6 10
ATGTTACCAT
AAGTTACGAT
AACAAAGCAA
AAGTTACCTT
AAGTTACCAA
TACTTACCAA
Sample Output
TAAGATAC
7
ACGTACGTAA
6
AAGTTACCAA
12
这个问题比较简单,注意字典序就好了。
附上AC代码:
#include<iostream>
#include<algorithm> using namespace std;
char str[][];
char ch[];
int t[];
int n;
int a,b;
int Sum;
int Flag;
int main()
{
cin >> n;
while(n--)
{
Sum=;
cin >> a >> b;
for(int i=;i<a;i++)
cin >> str[i];
for(int i=;i<b;i++)
{
Flag=;
t[]=t[]=t[]=t[]=;
for(int j=;j<a;j++)
{
if(str[j][i]=='A') t[]++;
if(str[j][i]=='C') t[]++;
if(str[j][i]=='G') t[]++;
if(str[j][i]=='T') t[]++;
}
for(int i=;i<;i++)
if(t[i]>t[Flag]) Flag=i;
if(Flag==) cout << 'A';
if(Flag==) cout << 'C';
if(Flag==) cout << 'G';
if(Flag==) cout << 'T';
Sum+=t[Flag];
}
cout << endl;
cout << a*b-Sum << endl;
}
return ;
}
uva1368 DNA Consensus String的更多相关文章
- UVA-1368 DNA Consensus String(思路)
题目: 链接 题意: 题目虽然比较长,但读完之后题目的思路还是比较容易想出来的. 给出m个长度为n的字符串(只包含‘A’.‘T’.‘G’.‘C’),我们的任务是得出一个字符串,要求这个字符串与给出的m ...
- 贪心水题。UVA 11636 Hello World,LA 3602 DNA Consensus String,UVA 10970 Big Chocolate,UVA 10340 All in All,UVA 11039 Building Designing
UVA 11636 Hello World 二的幂答案就是二进制长度减1,不是二的幂答案就是是二进制长度. #include<cstdio> int main() { ; ){ ; ) r ...
- DNA Consensus String
题目(中英对照): DNA (Deoxyribonucleic Acid) is the molecule which contains the genetic instructions. It co ...
- 紫书第三章训练1 E - DNA Consensus String
DNA (Deoxyribonucleic Acid) is the molecule which contains the genetic instructions. It consists of ...
- UVa1368/ZOJ3132 DNA Consensus String
#include <stdio.h>#include <string.h> int main(){ int a[4][1000]; // A/C/G/T在每列中出现的次数 ...
- DNA序列 (DNA Consensus String,ACM/ICPC Seoul 2006,UVa1368
题目描述:算法竞赛入门经典习题3-7 题目思路:每列出现最多的距离即最短 #include <stdio.h> #include <string.h> int main(int ...
- uva 1368 DNA Consensus String
这道题挺简单的,刚开始理解错误,以为是从已有的字符串里面求最短的距离,后面才发现是求一个到所有字符串最小距离的字符串,因为这样的字符串可能有多个,所以最后取最小字典序的字符串. 我的思路就是求每一列每 ...
- uvalive 3602 DNA Consensus String
https://vjudge.net/problem/UVALive-3602 题意: 给定m个长度均为n的DNA序列,求一个DNA序列,使得它到所有的DNA序列的汉明距离最短,若有多个解则输出字典序 ...
- 【每日一题】UVA - 1368 DNA Consensus String 字符串+贪心+阅读题
https://cn.vjudge.net/problem/UVA-1368 二维的hamming距离算法: For binary strings a and b the Hamming distan ...
随机推荐
- 使用typedef语句定义数组类型
使用typedef语句定义数组类型 1. 一维数组类型的定义格式 typedef <元素类型关键字><数组类型名>[<常量表达式>]; 例如: (1) ty ...
- PHP 中filter_var的使用
filter_var() 函数通过指定的过滤器过滤变量. 如果成功,则返回已过滤的数据,如果失败,则返回 false. 语法 :filter_var(variable, filter, options ...
- 2015第19周三小众app
今天搜集下个人知道好玩的小众app: 1.开眼每日推荐5个小视频,世界那么大,一起来看看吧. 2.懒人周末周末去哪了?简洁美观 3.LOFTER 网易良心出品. 4.玩儿去 发现很多很多好玩的去处 5 ...
- 定时关机命令-shutdown
定时关机命令-shutdown 一般会用到的定时关机命令有两种: Shutdown -s -t 3600 1小时后自动关机(3600秒) at 12:00 Shutdown -s 12:00自动关闭计 ...
- javascript 典型闭包的用法
<body><input type="radio" id="radio1" name="readionGroup" /&g ...
- hdu1695:数论+容斥
题目大意: 求x属于[1,b]和 y属于[1,d]的 gcd(x,y)=k 的方案数 题解: 观察发现 gcd()=k 不好处理,想到将x=x/k,y=y/k 后 gcd(x,y)=1.. 即问题转化 ...
- QQ地图api里的 地址解析函数 看不懂 javascript_百度知道
QQ地图api里的 地址解析函数 看不懂 javascript_百度知道 QQ地图api里的 地址解析函数 看不懂 javascript 2011-09-18 12:18 匿名 ...
- hdu 5391 Zball in Tina Town(打表找规律)
问题描述 Tina Town 是一个善良友好的地方,这里的每一个人都互相关心. Tina有一个球,它的名字叫zball.zball很神奇,它会每天变大.在第一天的时候,它会变大11倍.在第二天的时候, ...
- python 标准库基础学习之开发工具部分1学习
#2个标准库模块放一起学习,这样减少占用地方和空间#标准库之compileall字节编译源文件import compileall,re,sys#作用是查找到python文件,并把它们编译成字节码表示, ...
- 布局神器:Flexbox
最近的工作内容大多是移动端网页的开发,百分比布局,Media Queries,Bootstrap等常规的响应式/自适应的开发技术皆一一试过,但觉以上都不够灵活,所以,一直再尝试寻求更加灵活的精确的移动 ...
