吴裕雄--天生自然 R语言开发学习:分类(续二)








#-----------------------------------------------------------------------------#
# R in Action (2nd ed): Chapter 17 #
# Classification #
# requires packaged rpart, party, randomForest, kernlab, rattle #
# install.packages(c("rpart", "party", "randomForest", "e1071", "rpart.plot") #
# install.packages(rattle, dependencies = c("Depends", "Suggests")) #
#-----------------------------------------------------------------------------# par(ask=TRUE) # Listing 17.1 - Prepare the breast cancer data
loc <- "http://archive.ics.uci.edu/ml/machine-learning-databases/"
ds <- "breast-cancer-wisconsin/breast-cancer-wisconsin.data"
url <- paste(loc, ds, sep="") breast <- read.table(url, sep=",", header=FALSE, na.strings="?")
names(breast) <- c("ID", "clumpThickness", "sizeUniformity",
"shapeUniformity", "maginalAdhesion",
"singleEpithelialCellSize", "bareNuclei",
"blandChromatin", "normalNucleoli", "mitosis", "class") df <- breast[-1]
df$class <- factor(df$class, levels=c(2,4),
labels=c("benign", "malignant")) set.seed(1234)
train <- sample(nrow(df), 0.7*nrow(df))
df.train <- df[train,]
df.validate <- df[-train,]
table(df.train$class)
table(df.validate$class) # Listing 17.2 - Logistic regression with glm()
fit.logit <- glm(class~., data=df.train, family=binomial())
summary(fit.logit)
prob <- predict(fit.logit, df.validate, type="response")
logit.pred <- factor(prob > .5, levels=c(FALSE, TRUE),
labels=c("benign", "malignant"))
logit.perf <- table(df.validate$class, logit.pred,
dnn=c("Actual", "Predicted"))
logit.perf # Listing 17.3 - Creating a classical decision tree with rpart()
library(rpart)
set.seed(1234)
dtree <- rpart(class ~ ., data=df.train, method="class",
parms=list(split="information"))
dtree$cptable
plotcp(dtree) dtree.pruned <- prune(dtree, cp=.0125) library(rpart.plot)
prp(dtree.pruned, type = 2, extra = 104,
fallen.leaves = TRUE, main="Decision Tree") dtree.pred <- predict(dtree.pruned, df.validate, type="class")
dtree.perf <- table(df.validate$class, dtree.pred,
dnn=c("Actual", "Predicted"))
dtree.perf # Listing 17.4 - Creating a conditional inference tree with ctree()
library(party)
fit.ctree <- ctree(class~., data=df.train)
plot(fit.ctree, main="Conditional Inference Tree") ctree.pred <- predict(fit.ctree, df.validate, type="response")
ctree.perf <- table(df.validate$class, ctree.pred,
dnn=c("Actual", "Predicted"))
ctree.perf # Listing 17.5 - Random forest
library(randomForest)
set.seed(1234)
fit.forest <- randomForest(class~., data=df.train,
na.action=na.roughfix,
importance=TRUE)
fit.forest
importance(fit.forest, type=2)
forest.pred <- predict(fit.forest, df.validate)
forest.perf <- table(df.validate$class, forest.pred,
dnn=c("Actual", "Predicted"))
forest.perf # Listing 17.6 - A support vector machine
library(e1071)
set.seed(1234)
fit.svm <- svm(class~., data=df.train)
fit.svm
svm.pred <- predict(fit.svm, na.omit(df.validate))
svm.perf <- table(na.omit(df.validate)$class,
svm.pred, dnn=c("Actual", "Predicted"))
svm.perf # Listing 17.7 Tuning an RBF support vector machine (this can take a while)
set.seed(1234)
tuned <- tune.svm(class~., data=df.train,
gamma=10^(-6:1),
cost=10^(-10:10))
tuned
fit.svm <- svm(class~., data=df.train, gamma=.01, cost=1)
svm.pred <- predict(fit.svm, na.omit(df.validate))
svm.perf <- table(na.omit(df.validate)$class,
svm.pred, dnn=c("Actual", "Predicted"))
svm.perf # Listing 17.8 Function for assessing binary classification accuracy
performance <- function(table, n=2){
if(!all(dim(table) == c(2,2)))
stop("Must be a 2 x 2 table")
tn = table[1,1]
fp = table[1,2]
fn = table[2,1]
tp = table[2,2]
sensitivity = tp/(tp+fn)
specificity = tn/(tn+fp)
ppp = tp/(tp+fp)
npp = tn/(tn+fn)
hitrate = (tp+tn)/(tp+tn+fp+fn)
result <- paste("Sensitivity = ", round(sensitivity, n) ,
"\nSpecificity = ", round(specificity, n),
"\nPositive Predictive Value = ", round(ppp, n),
"\nNegative Predictive Value = ", round(npp, n),
"\nAccuracy = ", round(hitrate, n), "\n", sep="")
cat(result)
} # Listing 17.9 - Performance of breast cancer data classifiers
performance(dtree.perf)
performance(ctree.perf)
performance(forest.perf)
performance(svm.perf) # Using Rattle Package for data mining loc <- "http://archive.ics.uci.edu/ml/machine-learning-databases/"
ds <- "pima-indians-diabetes/pima-indians-diabetes.data"
url <- paste(loc, ds, sep="")
diabetes <- read.table(url, sep=",", header=FALSE)
names(diabetes) <- c("npregant", "plasma", "bp", "triceps",
"insulin", "bmi", "pedigree", "age", "class")
diabetes$class <- factor(diabetes$class, levels=c(0,1),
labels=c("normal", "diabetic"))
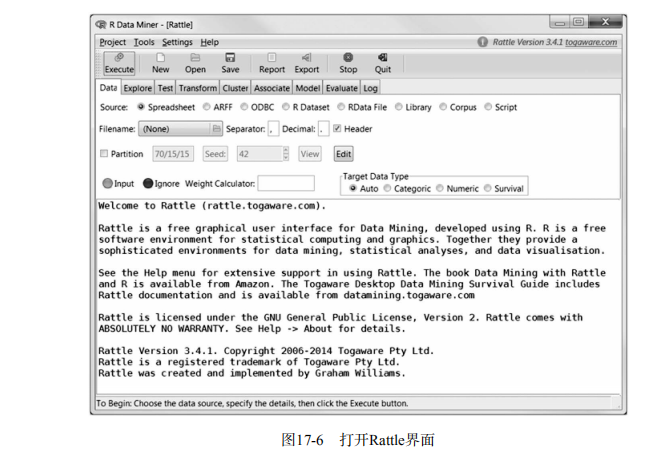
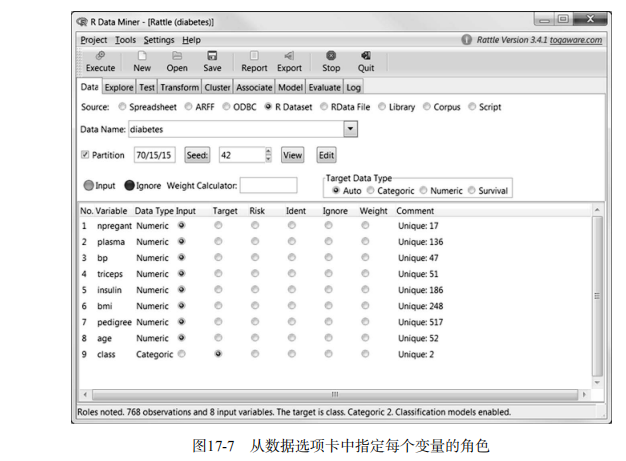
library(rattle)
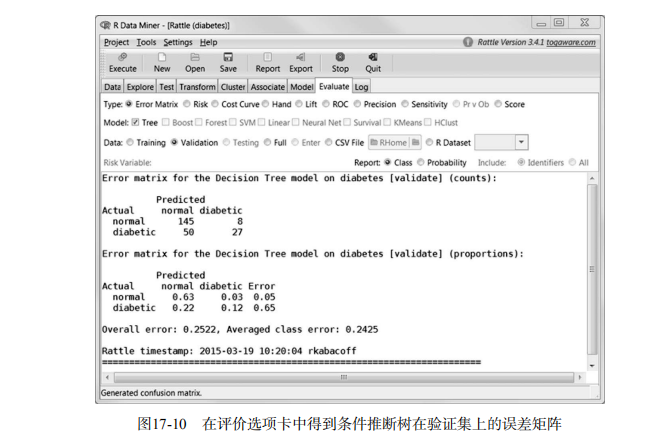
rattle()
吴裕雄--天生自然 R语言开发学习:分类(续二)的更多相关文章
- 吴裕雄--天生自然 R语言开发学习:R语言的安装与配置
下载R语言和开发工具RStudio安装包 先安装R
- 吴裕雄--天生自然 R语言开发学习:数据集和数据结构
数据集的概念 数据集通常是由数据构成的一个矩形数组,行表示观测,列表示变量.表2-1提供了一个假想的病例数据集. 不同的行业对于数据集的行和列叫法不同.统计学家称它们为观测(observation)和 ...
- 吴裕雄--天生自然 R语言开发学习:导入数据
2.3.6 导入 SPSS 数据 IBM SPSS数据集可以通过foreign包中的函数read.spss()导入到R中,也可以使用Hmisc 包中的spss.get()函数.函数spss.get() ...
- 吴裕雄--天生自然 R语言开发学习:使用键盘、带分隔符的文本文件输入数据
R可从键盘.文本文件.Microsoft Excel和Access.流行的统计软件.特殊格 式的文件.多种关系型数据库管理系统.专业数据库.网站和在线服务中导入数据. 使用键盘了.有两种常见的方式:用 ...
- 吴裕雄--天生自然 R语言开发学习:R语言的简单介绍和使用
假设我们正在研究生理发育问 题,并收集了10名婴儿在出生后一年内的月龄和体重数据(见表1-).我们感兴趣的是体重的分 布及体重和月龄的关系. 可以使用函数c()以向量的形式输入月龄和体重数据,此函 数 ...
- 吴裕雄--天生自然 R语言开发学习:基础知识
1.基础数据结构 1.1 向量 # 创建向量a a <- c(1,2,3) print(a) 1.2 矩阵 #创建矩阵 mymat <- matrix(c(1:10), nrow=2, n ...
- 吴裕雄--天生自然 R语言开发学习:图形初阶(续二)
# ----------------------------------------------------# # R in Action (2nd ed): Chapter 3 # # Gettin ...
- 吴裕雄--天生自然 R语言开发学习:图形初阶(续一)
# ----------------------------------------------------# # R in Action (2nd ed): Chapter 3 # # Gettin ...
- 吴裕雄--天生自然 R语言开发学习:图形初阶
# ----------------------------------------------------# # R in Action (2nd ed): Chapter 3 # # Gettin ...
- 吴裕雄--天生自然 R语言开发学习:基本图形(续二)
#---------------------------------------------------------------# # R in Action (2nd ed): Chapter 6 ...
随机推荐
- 题解 P4171 【[JSOI2010]满汉全席】
什么,tarjan?那是什么? 码量太大,我选择放弃 为什么不用dfs写2-sat呢?他会伤心的说 这题2-sat的过程大佬们已经讲得非常清楚了,我就略微提一下,主要讲dfs的原理 2_sat原理 我 ...
- 漫谈设计模式(一):代理(Proxy)模式与适配器(Adapter)模式对比
1.前言 为什么要将代理模式与适配器模式放在一起来说呢?因为它们有许多的共同点,当然也有一些不同的地方.首先两者都是属于结构型模式.结构型模型是这样定义的: 结构型模式涉及到如何组合类和类以获得更大的 ...
- Window RabbitMq安装
rabbitMQ是一个在AMQP协议标准基础上完整的,可服用的企业消息系统.它遵循Mozilla Public License开源协议,采用 Erlang 实现的工业级的消息队列(MQ)服务器,Rab ...
- CSP模拟赛游记
时间:2019.10.5 考试时间:100分钟(连正式考试时间的一半还没有到)题目:由于某些原因不能公开. 由于第一次接触NOIinux系统所以连怎么建文件夹,调字体,如何编译都不知道,考试的前半小时 ...
- Python之小作业
文档如下: # name, age, score tom, 12, 86 Lee, 15, 99 Lucy, 11, 58 Joseph, 19, 56 第一栏为姓名(name),第二栏为年纪(age ...
- ES6之模块化
本文介绍ES6实现模块化的方法:使用import和export. 导入的时候需不需要加大括号的判断:1.当用export default people导出时,就用 import people 导入(不 ...
- Docker容器化【Docker镜像与容器相关命令】
# Docker 学习目标: 掌握Docker基础知识,能够理解Docker镜像与容器的概念 完成Docker安装与启动 掌握Docker镜像与容器相关命令 掌握Tomcat Nginx 等软件的常用 ...
- 14 微服务电商【黑马乐优商城】:day02-springcloud(理论篇四:配置Robbin负载均衡)
本项目的笔记和资料的Download,请点击这一句话自行获取. day01-springboot(理论篇) :day01-springboot(实践篇) day02-springcloud(理论篇一) ...
- ios 获取app版本号
let infoDictionary = Bundle.main.infoDictionary!let appversion = infoDictionary["CFBundleShortV ...
- 记录几个windows常用的快捷键和命令
1.打开文件夹 win+E 2.关闭当前窗口 ctrl+w 3.切换窗口 alt+tab 4.输入命令窗口 win+r 5.注册表的快捷键 regedit 6.打开远程 mstsc 7.命令设置开机启 ...
